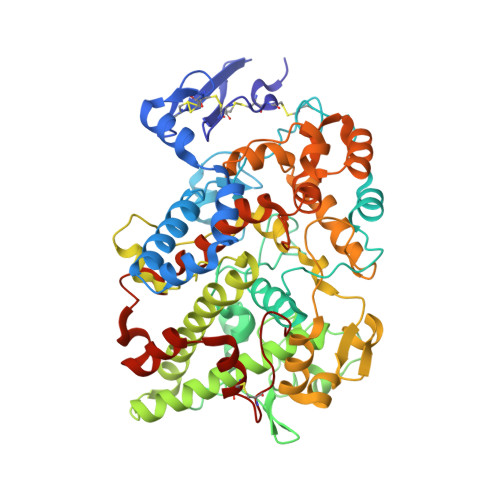
The 2.0A Resolution Crystal Structure of Prostaglandin H(2) Synthase-1: Structural Insights into an Unusual Peroxidase
Gupta, K., Selinsky, B.S., Kaub, C.J., Katz, A.K., Loll, P.J.(2004) J Mol Biology 335: 503-518
- PubMed: 14672659 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.10.073
- Primary Citation Related Structures:
1Q4G - PubMed Abstract:
Prostaglandin H2 synthase (EC 1.14.99.1) is an integral membrane enzyme containing a cyclooxygenase site, which is the target for the non-steroidal anti-inflammatory drugs, and a spatially distinct peroxidase site. Previous crystallographic studies of this clinically important drug target have been hindered by low resolution. We present here the 2.0 A resolution X-ray crystal structure of ovine prostaglandin H2 synthase-1 in complex with alpha-methyl-4-biphenylacetic acid, a defluorinated analog of the non-steroidal anti-inflammatory drug flurbiprofen. Detergent molecules are seen to bind to the protein's membrane-binding domain, and their positions suggest the depth to which this domain is likely to penetrate into the lipid bilayer. The relation of the enzyme's proximal heme ligand His388 to the heme iron is atypical for a peroxidase; the iron-histidine bond is unusually long and a substantial tilt angle is observed between the heme and imidazole planes. A molecule of glycerol, used as a cryoprotectant during diffraction experiments, is seen to bind in the peroxidase site, offering the first view of any ligand in this active site. Insights gained from glycerol binding may prove useful in the design of a peroxidase-specific ligand.
- Department of Biochemistry, Drexel University College of Medicine, 245 N 15th Street, Mailstop 497, Philadelphia, PA 19102-1192, USA.
Organizational Affiliation: