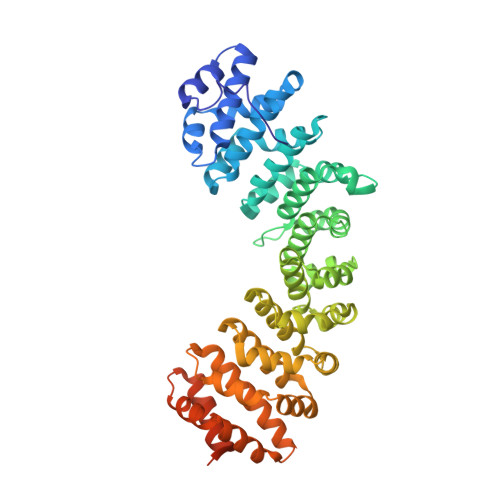Role of flanking sequences and phosphorylation in the recognition of the simian-virus-40 large T-antigen nuclear localization sequences by importin-alpha
Fontes, M.R.M., Teh, T., Toth, G., John, A., Pavo, I., Jans, D.A., Kobe, B.(2003) Biochem J 375: 339-349
- PubMed: 12852786 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20030510
- Primary Citation Related Structures:
1Q1S, 1Q1T - PubMed Abstract:
The nuclear import of simian-virus-40 large T-antigen (tumour antigen) is enhanced via phosphorylation by the protein kinase CK2 at Ser112 in the vicinity of the NLS (nuclear localization sequence). To determine the structural basis of the effect of the sequences flanking the basic cluster KKKRK, and the effect of phosphorylation on the recognition of the NLS by the nuclear import factor importin-alpha (Impalpha), we co-crystallized non-autoinhibited Impalpha with peptides corresponding to the phosphorylated and non-phosphorylated forms of the NLS, and determined the crystal structures of the complexes. The structures show that the amino acids N-terminally flanking the basic cluster make specific contacts with the receptor that are distinct from the interactions between bipartite NLSs and Impalpha. We confirm the important role of flanking sequences using binding assays. Unexpectedly, the regions of the peptides containing the phosphorylation site do not make specific contacts with the receptor. Binding assays confirm that phosphorylation does not increase the affinity of the T-antigen NLS to Impalpha. We conclude that the sequences flanking the basic clusters in NLSs play a crucial role in nuclear import by modulating the recognition of the NLS by Impalpha, whereas phosphorylation of the T-antigen enhances nuclear import by a mechanism that does not involve a direct interaction of the phosphorylated residue with Impalpha.
- Structural Biology Laboratory, St. Vincent's Institute of Medical Research, 41 Victoria Parade, Fitzroy, Victoria 3065, Australia.
Organizational Affiliation: