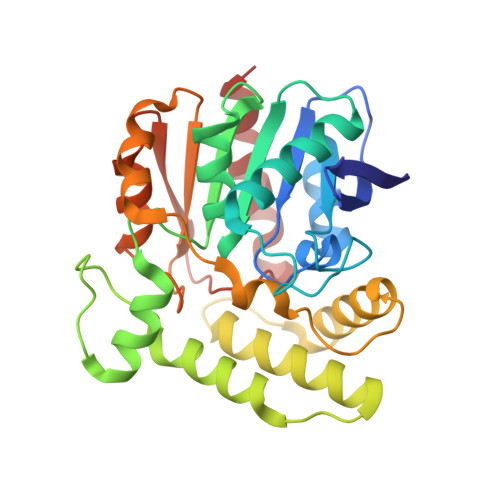
Crystal structure of aclacinomycin methylesterase with bound product analogues: implications for anthracycline recognition and mechanism.
Jansson, A., Niemi, J., Mantsala, P., Schneider, G.(2003) J Biological Chem 278: 39006-39013
- PubMed: 12878604 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304008200
- Primary Citation Related Structures:
1Q0R, 1Q0Z - PubMed Abstract:
Aclacinomycin methylesterase (RdmC) is one of the tailoring enzymes that modify the aklavinone skeleton in the biosynthesis of anthracyclines in Streptomyces species. The crystal structures of this enzyme from Streptomyces purpurascens in complex with the product analogues 10-decarboxymethylaclacinomycin T and 10-decarboxymethylaclacinomycin A were determined to nominal resolutions of 1.45 and 1.95 A, respectively. RdmC is built up of two domains. The larger alpha/beta domain shows the common alpha/beta hydrolase fold, whereas the smaller domain is alpha-helical. The active site and substrate binding pocket are located at the interface between the two domains. Decarboxymethylaclacinomycin T and decarboxymethylaclacinomycin A bind close to the catalytic triad (Ser102-His276-Asp248) in a hydrophobic pocket, with the sugar moieties located at the surface of the enzyme. The binding of the ligands is dominated by hydrophobic interactions, and specificity appears to be controlled mainly by the shape of the binding pocket rather than through specific hydrogen bonds. Mechanistic key features consistent with the structure of complexes of RdmC with product analogues are Ser102 acting as nucleophile and transition state stabilization by an oxyanion hole formed by the backbone amides of residues Gly32 and Met103.
- Molecular Structural Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, SE-171 77 Stockholm, Sweden.
Organizational Affiliation: