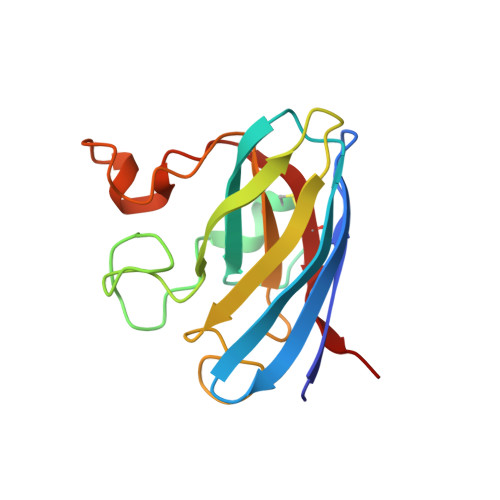
Structure of fully reduced bovine copper zinc superoxide dismutase at 1.15 A.
Hough, M.A., Hasnain, S.S.(2003) Structure 11: 937-946
- PubMed: 12906825 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00155-2
- Primary Citation Related Structures:
1Q0E - PubMed Abstract:
Copper zinc superoxide dismutase (CuZnSOD) forms a crucial component of the cellular response to oxidative stress by catalyzing the dismutation of the superoxide radical to hydrogen peroxide and water. Mutations in human CuZnSOD are associated with the development of familial amyotrophic lateral sclerosis (motor neuron disease). We have determined the structure of fully reduced bovine CuZnSOD to 1.15 A, the only atomic resolution structure for an intact CuZnSOD and one of only a small number for metalloproteins. For the first time, both subunits have been captured with the three coordinate Cu(I) ligation required by the generally accepted catalytic mechanism, where dismutation of the superoxide radical occurs via reduction of Cu. Furthermore, the improved resolution compared to previous studies (to 1.65 A) has allowed a more detailed examination of the metal center environment and its associated water network in the active site channel, facilitating the analysis of potential proton transfer routes.
- Molecular Biophysics Group, CCLRC Daresbury Laboratory, Daresbury, Warrington, Cheshire, WA4 4AD, United Kingdom.
Organizational Affiliation: