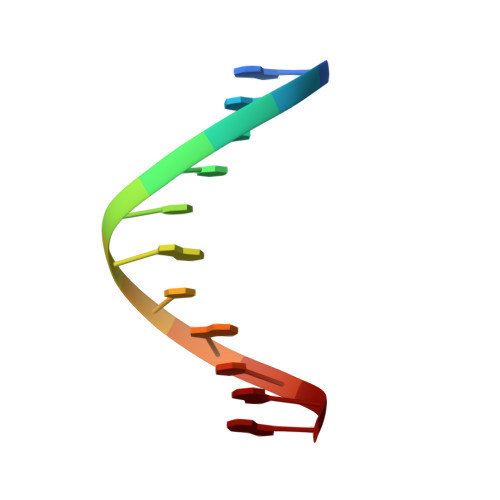
Solution structure of an O6-[4-oxo-4-(3-pyridyl)butyl]guanine adduct in an 11 mer DNA duplex: evidence for formation of a base triplex.
Peterson, L.A., Vu, C., Hingerty, B.E., Broyde, S., Cosman, M.(2003) Biochemistry 42: 13134-13144
- PubMed: 14609323
- DOI: https://doi.org/10.1021/bi035217v
- Primary Citation Related Structures:
1PYJ - PubMed Abstract:
The pyridyloxobutylating agents derived from metabolically activated tobacco-specific nitrosamines can covalently modify guanine bases in DNA at the O(6) position. The adduct formed, O(6)-[4-oxo-4-(3-pyridyl)butyl]guanine ([POB]dG), results in mutations that can lead to tumor formation, posing a significant cancer risk to humans exposed to tobacco smoke. A combined NMR-molecular mechanics computational approach was used to determine the solution structure of the [POB]dG adduct within an 11mer duplex sequence d(CCATAT-[POB]G-GCCC).d(GGGCCATATGG). In agreement with the NMR results, the POB ligand is located in the major groove, centered between the flanking 5'-side dT.dA and the 3'-side dG.dC base pairs and thus in the plane of the modified [POB]dG.dC base pair, which is displaced slightly into the minor groove. The modified base pair in the structure adopts wobble base pairing (hydrogen bonds between [POB]dG(N1) and dC(NH4) amino proton and between [POB]dG(NH2) amino proton and dC(N3)). A hydrogen bond appears to occur between the POB carbonyl oxygen and the partner dC's second amino proton. The modified guanine purine base, partner cytosine pyrimidine base, and POB pyridyl ring form a triplex via this unusual hydrogen-bonding pattern. The phosphodiester backbone twists at the lesion site, accounting for the unusual phosphorus chemical shift differences relative to those for the control DNA duplex. The helical distortions and wobble base pairing induced by the covalent binding of POB to the O(6)-position of dG help explain the significant decrease of 17.6 degrees C in melting temperature of the modified duplex relative to the unmodified control.
- Division of Environmental and Occupational Health and Cancer Center, University of Minnesota, Minneapolis, Minnesota 55455, USA.
Organizational Affiliation: