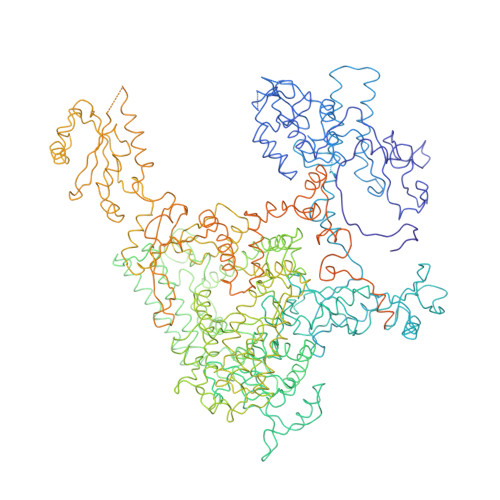
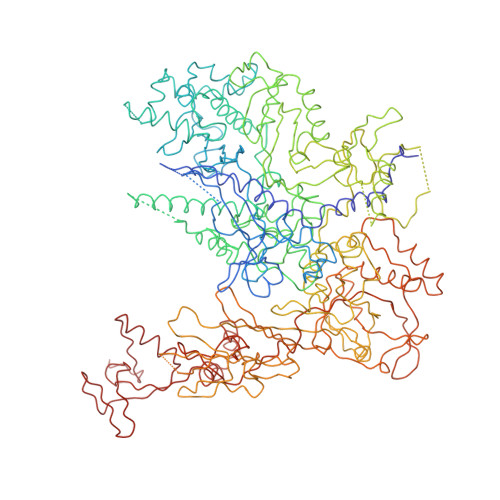
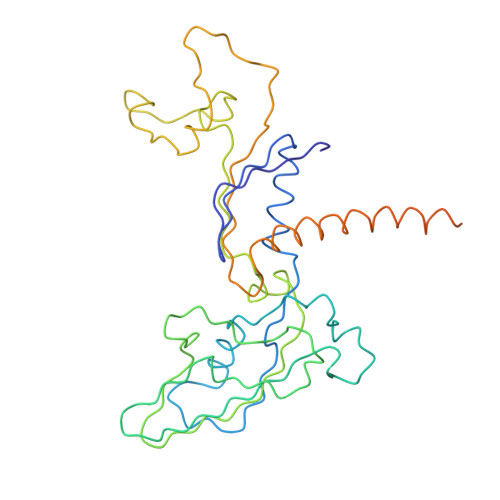
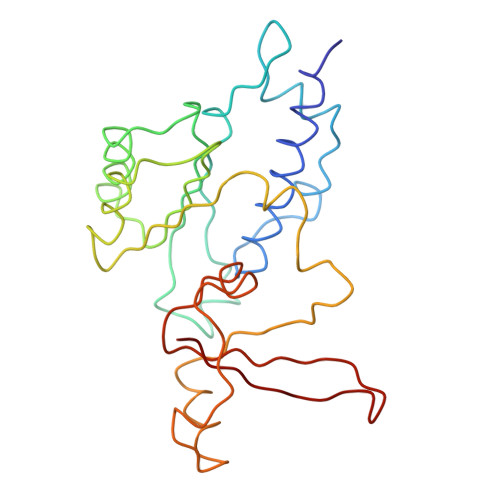
Architecture of the RNA Polymerase II-TFIIS Complex and Implications for mRNA Cleavage
Kettenberger, H., Armache, K.-J., Cramer, P.(2003) Cell 114: 347-357
- PubMed: 12914699 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(03)00598-1
- Primary Citation Related Structures:
1PQV - PubMed Abstract:
The transcription elongation factor TFIIS induces mRNA cleavage by enhancing the intrinsic nuclease activity of RNA polymerase (Pol) II. We have diffused TFIIS into Pol II crystals and derived a model of the Pol II-TFIIS complex from X-ray diffraction data to 3.8 A resolution. TFIIS extends from the polymerase surface via a pore to the internal active site, spanning a distance of 100 A. Two essential and invariant acidic residues in a TFIIS loop complement the Pol II active site and could position a metal ion and a water molecule for hydrolytic RNA cleavage. TFIIS also induces extensive structural changes in Pol II that would realign nucleic acids in the active center. Our results support the idea that Pol II contains a single tunable active site for RNA polymerization and cleavage, in contrast to DNA polymerases with two separate active sites for DNA polymerization and cleavage.
- Institute of Biochemistry, Gene Center, University of Munich, Feodor-Lynen-Str. 25, 81377 Munich, Germany.
Organizational Affiliation: