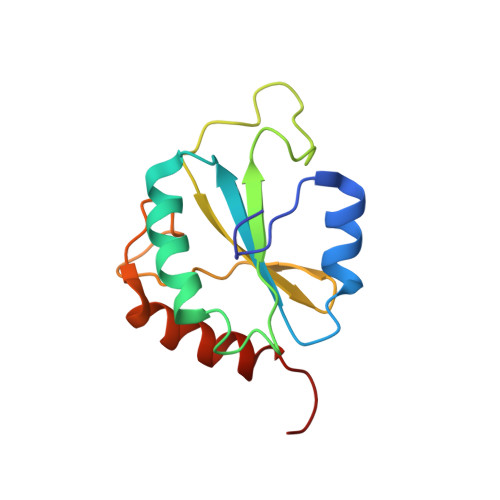
Structure, stability and function of hDim1 investigated by NMR, circular dichroism and mutational analysis
Zhang, Y.Z., Cheng, H., Gould, K.L., Golemis, E.A., Roder, H.(2003) Biochemistry 42: 9609-9618
- PubMed: 12911302
- DOI: https://doi.org/10.1021/bi034486i
- Primary Citation of Related Structures:
1PQN - PubMed Abstract:
The 142 amino acid Dim1p protein is a component of the U4/U6.U5 tri-snRNP complex required for pre-mRNA splicing and interacts with multiple splicing-associated proteins. To gain further insight into the structural basis of its function, we determined the solution structure of the reduced form of the dominant negative human hDim1 (hDim1(1)(-)(128)) using multidimensional NMR spectroscopy. This dominant negative hDim1 assumes a thioredoxin-like fold, confirming previous NMR and crystallographic results. However, in contrast to a recent crystal structure, the NMR solution structure for the reduced form of hDim1(1)(-)(128) presented here, along with thermodynamic data, indicates that the presence of a disulfide bond between Cys38 and Cys79 is structurally and functionally unimportant. Comparison of the truncated hDim1(1)(-)(128) with the full-length protein, using NMR and circular dichroism spectroscopy, indicates that the 14 C-terminal residues can undergo a local unfolding transition and assume alternative conformations, which appear to play a functional role. Other residues essential for hDim1 function are identified by using mutational and genetic approaches. The residues thus identified are not identical with those previously shown to govern Dim1 interaction with defined protein partners.
- Division of Basic Science, Fox Chase Cancer Center, Philadelphia, Pennsylvania 19111, USA. zhangy@iit.edu
Organizational Affiliation: