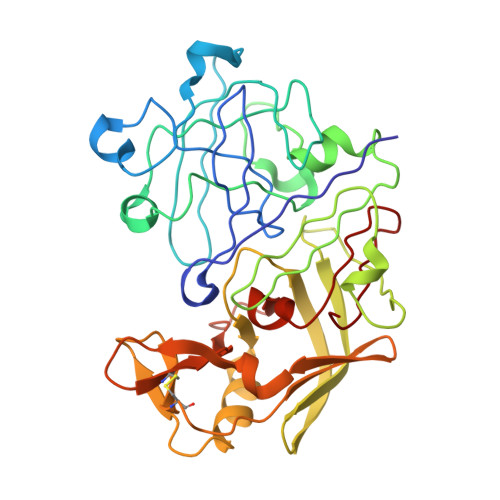
Crystallographic analysis of transition-state mimics bound to penicillopepsin: phosphorus-containing peptide analogues.
Fraser, M.E., Strynadka, N.C., Bartlett, P.A., Hanson, J.E., James, M.N.(1992) Biochemistry 31: 5201-5214
- PubMed: 1606144 Search on PubMed
- DOI: https://doi.org/10.1021/bi00137a016
- Primary Citation Related Structures:
1PPK, 1PPL, 1PPM - PubMed Abstract:
The molecular structures of three phosphorus-based peptide inhibitors of aspartyl proteinases complexed with penicillopepsin [1, Iva-L-Val-L-Val-StaPOEt [Iva = isovaleryl, StaP = the phosphinic acid analogue of statine [(S)-4-amino-(S)-3-hydroxy-6-methylheptanoic acid] (IvaVVStaPOEt)]; 2, Iva-L-Val-L-Val-L-LeuP-(O)Phe-OMe [LeuP = the phosphinic acid analogue of L-leucine; (O)Phe = L-3-phenyllactic acid; OMe = methyl ester] [Iva VVLP(O)FOMe]; and 3, Cbz-L-Ala-L-Ala-L-LeuP-(O)-Phe-OMe (Cbz = benzyloxycarbonyl) [CbzAALP(O)FOMe]] have been determined by X-ray crystallography and refined to crystallographic agreement factors, R ( = sigma parallel to F0 magnitude of - Fc parallel to/sigma magnitude of F0), of 0.132, 0.131, and 0.134, respectively. These inhibitors were designed to be structural mimics of the tetrahederal transition-state intermediate encountered during aspartic proteinase catalysis. They are potent inhibitors of penicillopepsin with Ki values of 1, 22 nM; 2, 2.8 nM; and 3, 1600 nM, respectively [Bartlett, P. A., Hanson, J. E., & Giannousis, P. P. (1990) J. Org. Chem. 55, 6268-6274]. All three of these phosphorus-based inhibitors bind virtually identically in the active site of penicillopepsin in a manner that closely approximates that expected for the transition state [James, M. N. G., Sielecki, A.R., Hayakawa, K., & Gelb, M. H. (1992) Biochemistry 31, 3872-3886]. The pro-S oxygen atom of the two phosphonate inhibitors and of the phosphinate group of the StaP inhibitor make very short contact distances (approximately 2.4 A) to the carboxyl oxygen atom, O delta 1, of Asp33 on penicillopepsin. We have interpreted this distance and the stereochemical environment of the carboxyl and phosphonate groups in terms of a hydrogen bond that most probably has a symmetric single-well potential energy function. The pro-R oxygen atom is the recipient of a hydrogen bond from the carboxyl group of Asp213. Thus, we are able to assign a neutral status to Asp213 and a partially negatively charged status to Asp33 with reasonable confidence. Similar very short hydrogen bonds involving the active site glutamic acid residues of thermolysin and carboxypeptidase A and the pro-R oxygen of bound phosphonate inhibitors have been reported [Holden, H. M., Tronrud, D. E., Monzingo, A. F., Weaver, L. H., & Matthews, B. W. (1987) Biochemistry 26, 8542-8553; Kim, H., & Lipscomb, W. N. (1991) Biochemistry 30, 8171-8180].(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Biochemistry, University of Alberta, Edmonton, Canada.
Organizational Affiliation: