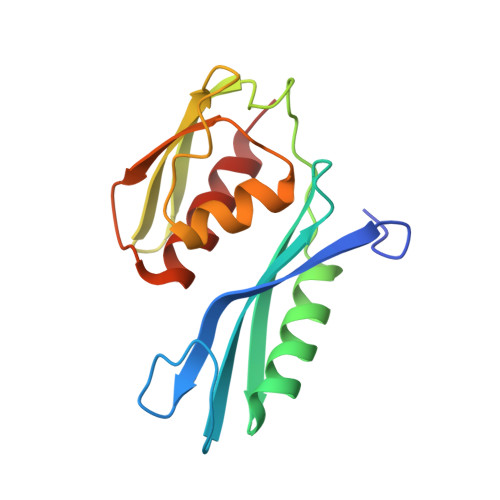
The structure of ribosomal protein S5 reveals sites of interaction with 16S rRNA.
Ramakrishnan, V., White, S.W.(1992) Nature 358: 768-771
- PubMed: 1508272
- DOI: https://doi.org/10.1038/358768a0
- Primary Citation of Related Structures:
1PKP - PubMed Abstract:
Understanding the process whereby the ribosome translates the genetic code into protein molecules will ultimately require high-resolution structural information, and we report here the first crystal structure of a protein from the small ribosomal subunit. This protein, S5, has a molecular mass of 17,500 and is highly conserved in all lifeforms. The molecule contains two distinct alpha/beta domains that have structural similarities to several other proteins that are components of ribonucleoprotein complexes. Mutations in S5 result in several phenotypes which suggest that S5 may have a role in translational fidelity and translocation. These include ribosome ambiguity or ram, reversion from streptomycin dependence and resistance to spectinomycin. Also, a cold-sensitive, spectinomycin-resistant mutant of S5 has been identified which is defective in initiation. Here we show that these mutations map to two distinct regions of the molecule which seem to be sites of interaction with ribosomal RNA. A structure/function analysis of the molecule reveals discrepancies with current models of the 30S subunit.
- Biology Department, Brookhaven National Laboratory, Upton, New York 11973.
Organizational Affiliation: