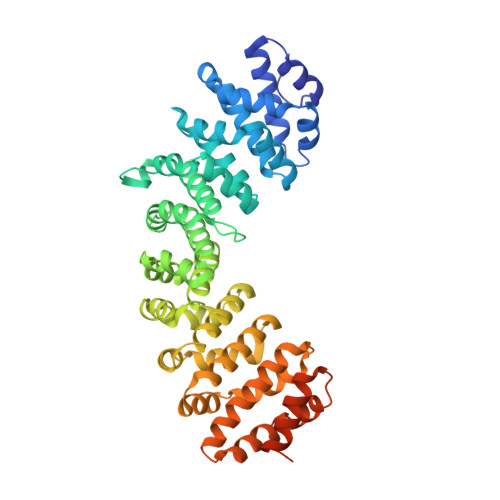
Structural basis for the specificity of bipartite nuclear localization sequence binding by importin-alpha
Fontes, M.R.M., Teh, T., Jans, D., Brinkworth, R.I., Kobe, B.(2003) J Biological Chem 278: 27981-27987
- PubMed: 12695505 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M303275200
- Primary Citation Related Structures:
1PJM, 1PJN - PubMed Abstract:
Importin-alpha is the nuclear import receptor that recognizes cargo proteins carrying conventional basic monopartite and bipartite nuclear localization sequences (NLSs) and facilitates their transport into the nucleus. Bipartite NLSs contain two clusters of basic residues, connected by linkers of variable lengths. To determine the structural basis of the recognition of diverse bipartite NLSs by mammalian importin-alpha, we co-crystallized a non-autoinhibited mouse receptor protein with peptides corresponding to the NLSs from human retinoblastoma protein and Xenopus laevis phosphoprotein N1N2, containing diverse sequences and lengths of the linker. We show that the basic clusters interact analogously in both NLSs, but the linker sequences adopt different conformations, whereas both make specific contacts with the receptor. The available data allow us to draw general conclusions about the specificity of NLS binding by importin-alpha and facilitate an improved definition of the consensus sequence of a conventional basic/bipartite NLS (KRX10-12KRRK) that can be used to identify novel nuclear proteins.
- Structural Biology Laboratory, St. Vincent's Institute of Medical Research, 41 Victoria Parade, Fitzroy, Victoria 3065, Australia.
Organizational Affiliation: