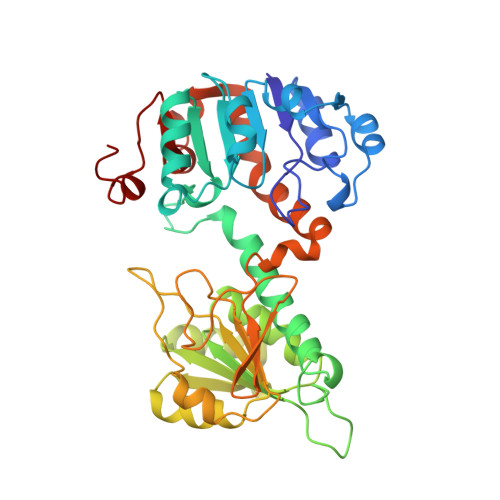
Analysis of the structure and substrate binding of Phormidium lapideum alanine dehydrogenase.
Baker, P.J., Sawa, Y., Shibata, H., Sedelnikova, S.E., Rice, D.W.(1998) Nat Struct Biol 5: 561-567
- PubMed: 9665169 Search on PubMed
- DOI: https://doi.org/10.1038/817
- Primary Citation Related Structures:
1PJB, 1PJC, 1SAY - PubMed Abstract:
The structure of the hexameric L-alanine dehydrogenase from Phormidium lapideum reveals that the subunit is constructed from two domains, each having the common dinucleotide binding fold. Despite there being no sequence similarity, the fold of alanine dehydrogenase is closely related to that of the family of D-2-hydroxyacid dehydrogenases, with a similar location of the active site, suggesting that these enzymes are related by divergent evolution. L-alanine dehydrogenase and the 2-hydroxyacid dehydrogenases also use equivalent functional groups to promote substrate recognition and catalysis. However, they are arranged differently on the enzyme surface, which has the effect of directing opposite faces of the keto acid to the dinucleotide in each case, forcing a change in absolute configuration of the product.
- The Krebs Institute, The Department of Molecular Biology & Biotechnology, The University of Sheffield, UK. P.baker@sheffield.ac.uk
Organizational Affiliation: