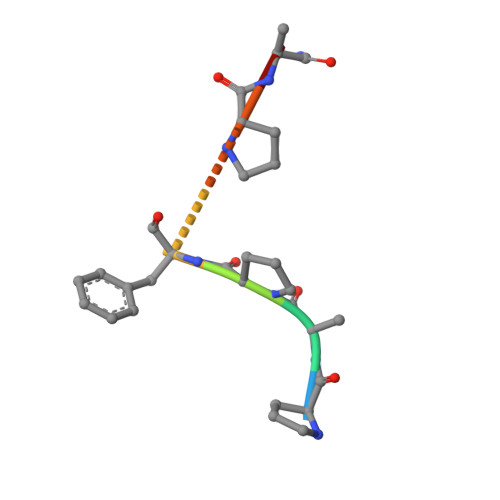
Structure of a ternary complex of proteinase K, mercury, and a substrate-analogue hexa-peptide at 2.2 A resolution
Saxena, A.K., Singh, T.P., Peters, K., Fittkau, S., Visanji, M., Wilson, K.S., Betzel, C.(1996) Proteins 25: 195-201
- PubMed: 8811735 Search on PubMed
- DOI: https://doi.org/10.1002/(SICI)1097-0134(199606)25:2<195::AID-PROT5>3.0.CO;2-H
- Primary Citation Related Structures:
1PJ8 - PubMed Abstract:
The crystal structure of a ternary complex of proteinase K, Hg(II) and a hexapeptide N-Ac-Pro-Ala-Pro-Phe-Pro-Ala-NH2 has been determined at 2.2 A resolution and refined to an R factor of 0.172 for 12,910 reflections. The mercury atom occupies two alternate sites, each of which was assigned an occupancy of 0.45. These two sites are bridged by Cys-73 S gamma which forms covalent bonds to both. Both mercury sites form regular polyhedrons involving atoms from residues Asp-39, His-69, Cys-73, His-72, Met-225, and Wat-324. The complex formation with mercury seems to disturb the stereochemistry of the residues of the catalytic triad Asp-39, His-69, and Ser-224 appreciably, thus reducing the enzymatic activity of proteinase K to 15%. The electron density in the difference Fourier map shows that the hexapeptide occupies the S1 subsite predominantly and the standard recognition site constituted by Ser-132 to Gly-136 and Gly-100 to Tyr-104 segments is virtually empty. The hexapeptide is held firmly through a series of hydrogen bonds involving protein atoms and water molecules. As a result of complex formation, Asp-39, His-69, Met-225, Ile-220, Ser-219, Thr-223, and Ser-224 residues move appreciably to accommodate the mercury atoms and the hexapeptide. The largest movement is observed for Met-225 which is involved in multiple interactions with both mercury and the hexapeptide. The activity results indicate an inhibition rate of 95%, as a result of the combined effect of mercury and hexapeptide.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi, India.
Organizational Affiliation: