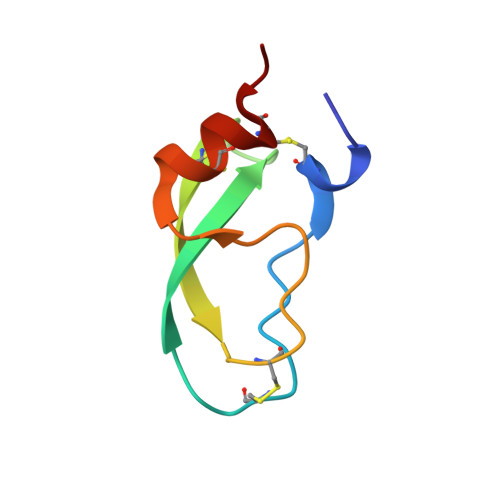
Determination of a high-quality nuclear magnetic resonance solution structure of the bovine pancreatic trypsin inhibitor and comparison with three crystal structures.
Berndt, K.D., Guntert, P., Orbons, L.P., Wuthrich, K.(1992) J Mol Biology 227: 757-775
- PubMed: 1383552 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90222-6
- Primary Citation Related Structures:
1PIT - PubMed Abstract:
A high-quality three-dimensional structure of the bovine pancreatic trypsin inhibitor (BPTI) in aqueous solution was determined by 1H nuclear magnetic resonance (n.m.r.) spectroscopy and compared to the three available high-resolution X-ray crystal structures. A newly collected input of 642 distance constraints derived from nuclear Overhauser effects and 115 dihedral angle constraints was used for the structure calculations with the program DIANA, followed by restrained energy minimization with the program AMBER. The BPTI solution structure is represented by a group of 20 conformers with an average root-mean-square deviation (RMSD) relative to the mean solution structure of 0.43 A for backbone atoms and 0.92 A for all heavy atoms of residues 2 to 56. The pairwise RMSD values of the three crystal structures relative to the mean solution structure are 0.76 to 0.85 A for the backbone atoms and 1.24 to 1.33 A for all heavy atoms of residues 2 to 56. Small local differences in backbone atom positions between the solution structure and the X-ray structures near residues 9, 25 to 27, 46 to 48 and 52 to 58, and conformational differences for individual amino acid side-chains were analyzed for possible correlations with intermolecular protein-protein contacts in the crystal lattices, using the pairwise RMSD values among the three crystal structures as a reference.
- Institut für Molekularbiologie und Biophysik, Eidgenösische Technische Hochschule-Hönggerberg, Zürich, Switzerland.
Organizational Affiliation: