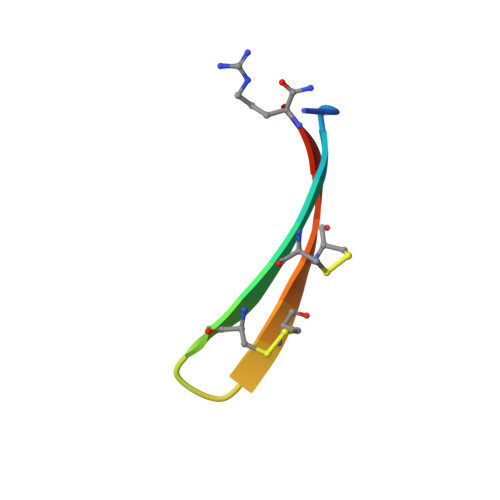
Solution structure of protegrin-1, a broad-spectrum antimicrobial peptide from porcine leukocytes.
Fahrner, R.L., Dieckmann, T., Harwig, S.S., Lehrer, R.I., Eisenberg, D., Feigon, J.(1996) Chem Biol 3: 543-550
- PubMed: 8807886
- DOI: https://doi.org/10.1016/s1074-5521(96)90145-3
- Primary Citation Related Structures:
1PG1 - PubMed Abstract:
The protegrins are a family of arginine- and cysteine-rich cationic peptides found in porcine leukocytes that exhibit a broad range of antimicrobial and antiviral activities. They are composed of 16-18 amino-acid residues including four cysteines, which form two disulfide linkages. To begin to understand the mechanism of action of these peptides, we set out to determine the structure of protegrin-1 (PG-1). We used two-dimensional homonuclear nuclear magnetic resonance spectroscopy to study the conformation of both natural and synthetic PG-1 under several conditions. A refined three-dimensional structure of synthetic PG-1 is presented. Both synthetic and natural protegrin-1 form a well-defined structure in solution composed primarily of a two-stranded antiparallel beta sheet, with strands connected by a beta turn. The structure of PG-1 suggests ways in which the peptide may interact with itself or other molecules to form the membrane pores and the large membrane-associated assemblages observed in protegrin-treated, gram-negative bacteria.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, CA 90095, USA. feigon@ewald.mbi.ucla.edu
Organizational Affiliation: