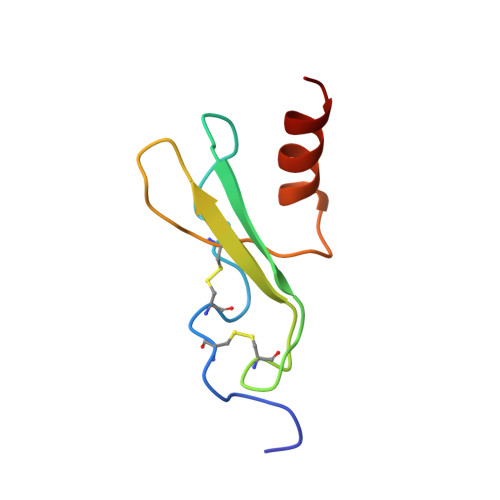
NMR solution structure of the 32-kDa platelet factor 4 ELR-motif N-terminal chimera: a symmetric tetramer.
Mayo, K.H., Roongta, V., Ilyina, E., Milius, R., Barker, S., Quinlan, C., La Rosa, G., Daly, T.J.(1995) Biochemistry 34: 11399-11409
- PubMed: 7547867 Search on PubMed
- DOI: https://doi.org/10.1021/bi00036a012
- Primary Citation Related Structures:
1PFM, 1PFN - PubMed Abstract:
Native human platelet factor 4 (PF4) is a homotetrameric protein (70 residues/subunit) known for its anticoagulant heparin binding activity. 2D 15N--1H HSQC NMR experiments of native PF4 in solution show the presence of conformational heterogeneity consistent with the formation of asymmetric homo-tetramers as observed in the X-ray crystal structure of both human and bovine PF4. A chimeric mutant of PF4 (called PF4-M2) which substitutes the first 11 N-terminal residues for the first eight residues from homologous interleukin-8 forms symmetric homo-tetramers with essentially the same heparin binding activity as native PF4. The solution structure of PF4-M2 has been investigated by using two- and three-dimensional 1H- and 15N-NMR spectroscopy and NOE-restrained simulated annealing molecular dynamics. As with other members of the CXC chemokine family whose structures are known, the PF4-M2 subunit monomer consists of a mostly hydrophobic, triple-stranded antiparallel beta-sheet onto which is folded an amphipathic C-terminal helix and a less periodic N-terminal domain. Although N-terminal substitution with the less acidic interleukin-8 sequence most affects the quarternary structure relative to native PF4 at the AC and AD dimer interfaces, AB dimer stability is weakened as reflected in reduced equilibrium association binding constants.
- Department of Biochemistry, University of Minnesota, Minneapolis 55455, USA.
Organizational Affiliation: