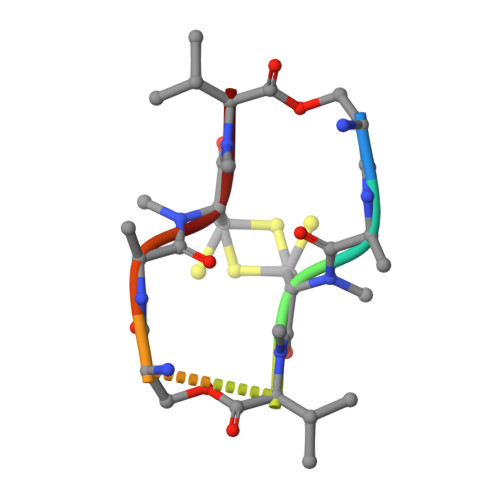
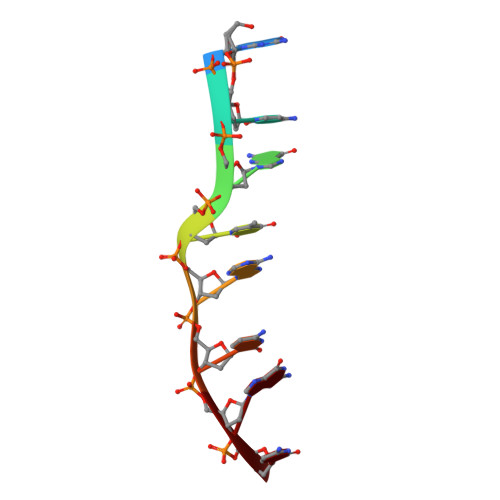
Structures of Complexes between Echinomycin and Duplex DNA.
Cuesta-Seijo, J.A., Sheldrick, G.M.(2005) Acta Crystallogr D Biol Crystallogr 61: 442
- PubMed: 15805599 Search on PubMed
- DOI: https://doi.org/10.1107/S090744490500137X
- Primary Citation Related Structures:
1PFE, 1XVK, 1XVN, 1XVR - PubMed Abstract:
The structure of the bis-intercalation complex of the depsipeptide antibiotic echinomycin with (CGTACG)2 has been redetermined at a higher resolution (1.4 A) and new high-resolution structures (1.1-1.5 A) are reported for the complexes of echinomycin with (GCGTACGC)2 (at both low and high ionic strengths) and (ACGTACGT)2. The structures show the expected Hoogsteen pairing for the base pairs flanking the intercalating chromophores on the outside and Watson-Crick pairing for both base pairs enclosed by the echinomycin. In the octamer complexes but not the hexamer complex, the echinomycin molecule, which would possess a molecular twofold axis were it not for the thioacetal bridge, shows twofold disorder. In all the structures the stacking of the base pairs and chromophores is extended by intermolecular stacking. The structures provide more precise details of the hydrogen bonding and other interactions between the bis-intercalating antibiotics and the duplex DNA than were previously available.
- Department of Structural Chemistry, University of Göttingen, Tammannstrasse 4, D37077 Göttingen, Germany.
Organizational Affiliation: