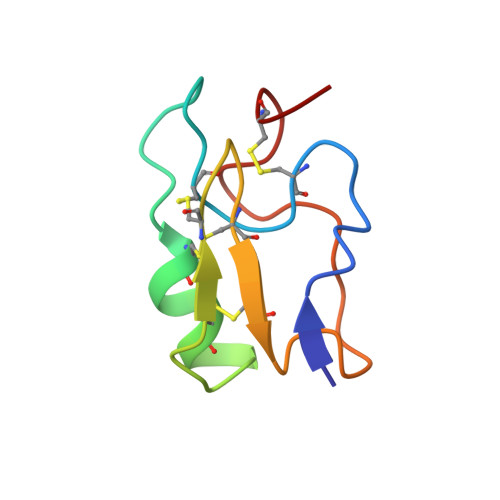
NMR solution structure of Cn12, a novel peptide from the Mexican scorpion Centruroides noxius with a typical beta-toxin sequence but with alpha-like physiological activity.
Del Rio-Portilla, F., Hernandez-Marin, E., Pimienta, G., Coronas, F.V., Zamudio, F.Z., Rodrguez de la Vega, R.C., Wanke, E., Possani, L.D.(2004) Eur J Biochem 271: 2504-2516
- PubMed: 15182366 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.2004.04181.x
- Primary Citation Related Structures:
1PE4 - PubMed Abstract:
Cn12 isolated from the venom of the scorpion Centruroides noxius has 67 amino-acid residues, closely packed with four disulfide bridges. Its primary structure and disulfide bridges were determined. Cn12 is not lethal to mammals and arthropods in vivo at doses up to 100 microg per animal. Its 3D structure was determined by proton NMR using 850 distance constraints, 36 phi angles derived from 36 coupling constants obtained by two different methods, and 22 hydrogen bonds. The overall structure has a two and half turn alpha-helix (residues 24-32), three strands of antiparallel beta-sheet (residues 2-4, 37-40 and 45-48), and a type II turn (residues 41-44). The amino-acid sequence of Cn12 resembles the beta scorpion toxin class, although patch-clamp experiments showed the induction of supplementary slow inactivation of Na(+) channels in F-11 cells (mouse neuroblastoma N18TG-2 x rat DRG2), which means that it behaves more like an alpha scorpion toxin. This behaviour prompted us to analyse Na(+) channel binding sites using information from 112 Na(+) channel gene clones available in the literature, focusing on the extracytoplasmic loops of the S5-S6 transmembrane segments of domain I and the S3-S4 segments of domain IV, sites considered to be responsible for binding alpha scorpion toxins.
- Institute of Chemistry, National Autonomous University of Mexico, Mexico City, Mexico.
Organizational Affiliation: