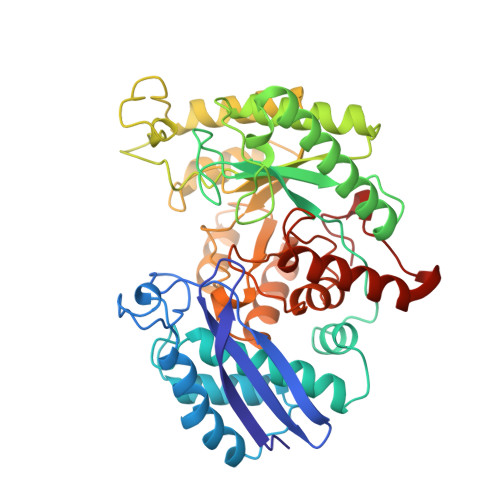
X-ray structure and catalytic mechanism of lobster enolase.
Duquerroy, S., Camus, C., Janin, J.(1995) Biochemistry 34: 12513-12523
- PubMed: 7547999 Search on PubMed
- Primary Citation Related Structures:
1PDY, 1PDZ - PubMed Abstract:
Enolase prepared from lobster tail muscle yielded trigonal crystals with one 47 kDa subunit per asymmetric unit. X-ray data were collected on the apoenzyme at 2.4 A resolution and on a complex with Mn2+ and the inhibitor phosphoglycolate at 2.2 A resolution. The corresponding cDNA was amplified from a library of lobster muscle cDNA, and a sequence corresponding to residues 27-398 was determined. It is highly homologous to other enolases, including yeast enolase for which an X-ray structure is available. Yeast enolase was used as a starting point for crystallographic refinement, which led to models of lobster enolase having R-factors below 22% and good stereochemistry. These models are very similar to yeast enolase; they have the same fold with a beta 3 alpha 4 N-terminal domain followed by an atypical alpha/beta barrel. Lobster apoenolase and the ternary complex differ only in the position of three mobile loops. In the complex, a single Mn2+ ion is seen ligated to three carboxylates and three water molecules. Phosphoglycolate binds near, but not directly to, the metal. His 157, which belongs to one of the mobile loops, is in contact with the C2 atom of the ligand. A water molecule hydrogen-bonds to the carboxylate of the ligand and to those of Glu 166 and Glu 209. We suggest that His 157 is the base that abstracts the C2H proton, whereas the water molecule is part of a proton relay system keeping the substrate in the carboxylic acid form where the pKa of the C2H group is low enough for proton transfer to His 157. The resulting catalytic mechanism is different from those proposed on the basis of the yeast enzyme X-ray structures, but it fits with earlier biochemical and spectroscopic data.
- Laboratoire de Biologie Structurale, UMR 9920 CNRS, Université Paris-Sud, Gif-sur-Yvette, France.
Organizational Affiliation: