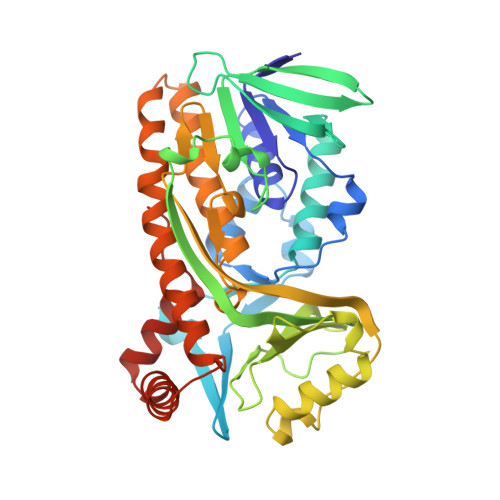
Crystal structure of p-hydroxybenzoate hydroxylase reconstituted with the modified FAD present in alcohol oxidase from methylotrophic yeasts: evidence for an arabinoflavin.
van Berkel, W.J., Eppink, M.H., Schreuder, H.A.(1994) Protein Sci 3: 2245-2253
- PubMed: 7756982 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560031210
- Primary Citation Related Structures:
1PDH - PubMed Abstract:
The flavin prosthetic group (FAD) of p-hydroxybenzoate hydroxylase from Pseudomonas fluorescens was replaced by a stereochemical analog, which is spontaneously formed from natural FAD in alcohol oxidases from methylotrophic yeasts. Reconstitution of p-hydroxybenzoate hydroxylase from apoprotein and modified FAD is a rapid process complete within seconds. Crystals of the enzyme-substrate complex of modified FAD-containing p-hydroxybenzoate hydroxylase diffract to 2.1 A resolution. The crystal structure provides direct evidence for the presence of an arabityl sugar chain in the modified form of FAD. The isoalloxazine ring of the arabinoflavin adenine dinucleotide (a-FAD) is located in a cleft outside the active site as recently observed in several other p-hydroxybenzoate hydroxylase complexes. Like the native enzyme, a-FAD-containing p-hydroxybenzoate hydroxylase preferentially binds the phenolate form of the substrate (pKo = 7.2). The substrate acts as an effector highly stimulating the rate of enzyme reduction by NADPH (kred > 500 s-1). The oxidative part of the catalytic cycle of a-FAD-containing p-hydroxybenzoate hydroxylase differs from native enzyme. Partial uncoupling of hydroxylation results in the formation of about 0.3 mol of 3,4-dihydroxybenzoate and 0.7 mol of hydrogen peroxide per mol NADPH oxidized. It is proposed that flavin motion in p-hydroxybenzoate hydroxylase is important for efficient reduction and that the flavin "out" conformation is associated with the oxidase activity.
- Department of Biochemistry, Agricultural University, Wageningen, The Netherlands.
Organizational Affiliation: