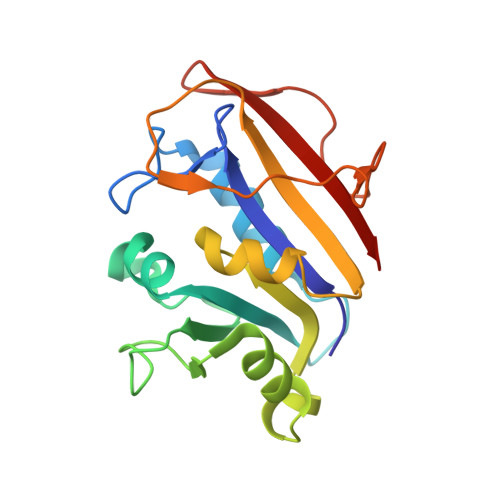
Analysis of three crystal structure determinations of a 5-methyl-6-N-methylanilino pyridopyrimidine antifolate complex with human dihydrofolate reductase.
Cody, V., Luft, J.R., Pangborn, W., Gangjee, A.(2003) Acta Crystallogr D Biol Crystallogr 59: 1603-1609
- PubMed: 12925791
- DOI: https://doi.org/10.1107/s0907444903014963
- Primary Citation of Related Structures:
1PD8, 1PD9, 1PDB - PubMed Abstract:
Structural data are reported for the first example of the potent antifolate inhibitor 2,4-diamino-5-methyl-6-[(3',4',5'-trimethoxy-N-methylanilino)methyl]pyrido[2,3-d]pyrimidine (1) in complex with human dihydrofolate reductase (hDHFR) and NADPH. Small differences in crystallization conditions resulted in the growth of two different forms of a binary complex. The structure determination of an additional crystal of a ternary complex of hDHFR with NADPH and (1) grown under similar conditions is also reported. Diffraction data were collected to 2.1 A resolution for an R3 lattice from a hDHFR ternary complex with NADPH and (1) and to 2.2 A resolution from a binary complex. Data were also collected to 2.1 A resolution from a binary complex with hDHFR and (1) in the first example of a tetragonal P4(3)2(1)2 lattice. Comparison of the intermolecular contacts among these structures reveals differences in the backbone conformation (1.9-3.2 A) for flexible loop regions (residues 40-46, 77-83 and 103-107) that reflect differences in the packing environment between the rhombohedral and tetragonal space groups. Analysis of the packing environments shows that the tetragonal lattice is more tightly packed, as reflected in its smaller V(M) value and lower solvent content. The conformation of the inhibitor (1) is similar in all structures and is also similar to that observed for TMQ, the parent quinazoline compound. The activity profile for this series of 5-deaza N-substituted non-classical trimethoxybenzyl antifolates shows that the N10-CH(3) substituted (1) has the greatest potency and selectivity for Toxoplasma gondii DHFR (tgDHFR) compared with its N-H or N-CHO analogs. Models of the tgDHFR active site indicate preferential contacts with (1) that are not present in either the human or Pneumocystis carinii DHFR structures. Differences in the acidic residue (Glu30 versus Asp for tgDHFR) affect the precise positioning of the diaminopyridopyrimidine ring, while changes in other residues, particularly at positions 60 and 64 (Leu versus Met and Asn versus Phe), involve interactions with the trimethoxybenzyl substituents.
- Structural Biology Department, Hauptman-Woodward Medical Research Institute Inc., 73 High Street, Buffalo, NY 14203, USA. cody@hwi.buffalo.edu
Organizational Affiliation: