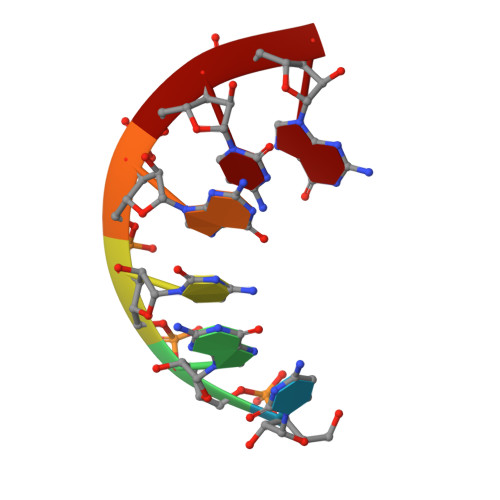
Solution structure of RNA duplexes containing alternating CG base pairs: NMR study of r(CGCGCG)2 and 2'-O-Me(CGCGCG)2 under low salt conditions.
Popenda, M., Biala, E., Milecki, J., Adamiak, R.W.(1997) Nucleic Acids Res 25: 4589-4598
- PubMed: 9358170 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/25.22.4589
- Primary Citation Related Structures:
1PBL, 1PBM - PubMed Abstract:
Structures of r(CGCGCG)2 and 2'-O-Me(CGCGCG)2 have been determined by NMR spectroscopy under low salt conditions. All protons and phosphorus nuclei resonances have been assigned. Signals of H5'/5" have been assigned stereospecifically. All 3JH,H and 3JP,H coupling constants have been measured. The structures were determined and refined using an iterative relaxation matrix procedure (IRMA) and the restrained MD simulation. Both duplexes form half-turn, right-handed helices with several conformational features which deviate significantly from a canonical A-RNA structure. Duplexes are characterised as having C3'-endo sugar pucker, very low base-pair rise and high helical twist and inclination angles. Helices are overwound with <10 bp per turn. There is limited inter-strand guanine stacking for CG steps. Within CG steps of both duplexes, the planes of the inter-strand cytosines are not parallel while guanines are almost parallel. For the GC steps this pattern is reversed. The 2'-O-methyl groups are spatially close to the 5'-hydrogens of neighbouring residues from the 3'-side and are directed towards the minor groove of 2'-O-Me(CGCGCG)2 forming a hydrophobic layer. Solution structures of both duplexes are similar; the effect of 2'-O-methylation on the parent RNA structure is small. This suggests that intrinsic properties imposed by alternating CG base pairs govern the overall conformation of both duplexes.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Noskowskiego 12/14, 61-704 Poznan, Poland.
Organizational Affiliation: