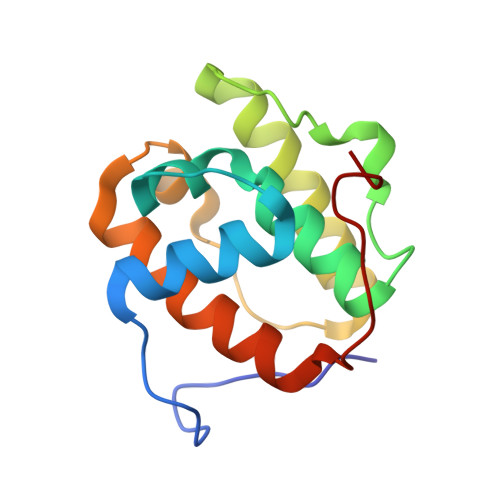
Crystal structure of the amino-terminal microtubule-binding domain of end-binding protein 1 (EB1)
Hayashi, I., Ikura, M.(2003) J Biological Chem 278: 36430-36434
- PubMed: 12857735
- DOI: https://doi.org/10.1074/jbc.M305773200
- Primary Citation of Related Structures:
1PA7, 1UEG - PubMed Abstract:
The end-binding protein 1 (EB1) family is a highly conserved group of proteins that localizes to the plus-ends of microtubules. EB1 has been shown to play an important role in regulating microtubule dynamics and chromosome segregation, but its regulation mechanism is poorly understood. We have determined the 1.45-A resolution crystal structure of the amino-terminal domain of EB1, which is essential for microtubule binding, and show that it forms a calponin homology (CH) domain fold that is found in many proteins involved in the actin cytoskeleton. The functional CH domain for actin binding is a tandem pair, whereas EB1 is the first example of a single CH domain that can associate with the microtubule filament. Although our biochemical study shows that microtubule binding of EB1 is electrostatic in part, our mutational analysis suggests that the hydrophobic network, which is partially exposed in our crystal structure, is also important for the association. We propose that, like other actin-binding CH domains, EB1 employs the hydrophobic interaction to bind to microtubules.
- Division of Molecular and Structural Biology, Ontario Cancer Institute and Department of Medical Biophysics, University of Toronto, Toronto, Ontario M5G 2M9, Canada. ihayashi@uhnres.utoronto.ca
Organizational Affiliation: