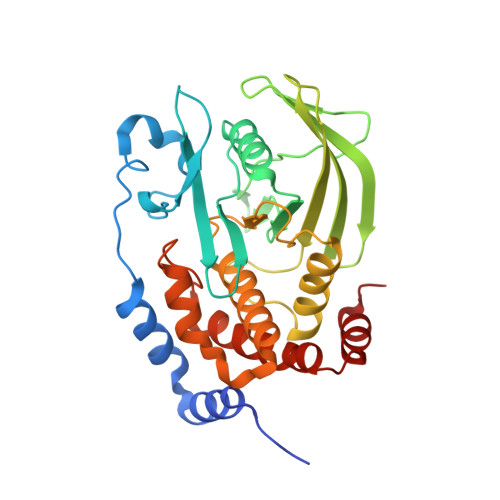
Functional characterization and crystal structure of the C215D mutant of protein-tyrosine phosphatase-1B
Romsicki, Y., Scapin, G., Beaulieu-Audy, V., Patel, S., Becker, J.W., Kennedy, B.P., Asante-Appiah, E.(2003) J Biological Chem 278: 29009-29015
- PubMed: 12748196 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M303817200
- Primary Citation Related Structures:
1PA1 - PubMed Abstract:
We have characterized the C215D active-site mutant of protein-tyrosine phosphatase-1B (PTP-1B) and solved the crystal structure of the catalytic domain of the apoenzyme to a resolution of 1.6 A. The mutant enzyme displayed maximal catalytic activity at pH approximately 4.5, which is significantly lower than the pH optimum of 6 for wild-type PTP-1B. Although both forms of the enzyme exhibited identical Km values for hydrolysis of p-nitrophenyl phosphate at pH 4.5 and 6, the kcat values of C215D were approximately 70- and approximately 7000-fold lower than those of wild-type PTP-1B, respectively. Arrhenius plots revealed that the mutant and wild-type enzymes displayed activation energies of 61 +/- 1 and 18 +/- 2 kJ/mol, respectively, at their pH optima. Unlike wild-type PTP-1B, C215D-mediated p-nitrophenyl phosphate hydrolysis was inactivated by 1,2-epoxy-3-(p-nitrophenoxy)propane, suggesting a direct involvement of Asp215 in catalysis. Increasing solvent microviscosity with sucrose (up to 40% (w/v)) caused a significant decrease in kcat/Km of the wild-type enzyme, but did not alter the catalytic efficiency of the mutant protein. Structurally, the apoenzyme was identical to wild-type PTP-1B, aside from the flexible WPD loop region, which was in both "open" and "closed" conformations. At physiological pH, the C215D mutant of PTP-1B should be an effective substrate-trapping mutant that can be used to identify cellular substrates of PTP-1B. In addition, because of its insensitivity to oxidation, this mutant may be used for screening fermentation broth and other natural products to identify inhibitors of PTP-1B.
- Department of Biochemistry and Molecular Biology, Merck Frosst Centre for Therapeutic Research, Pointe-Claire, Dorval, Quebec H9R 4P8, Canada.
Organizational Affiliation: