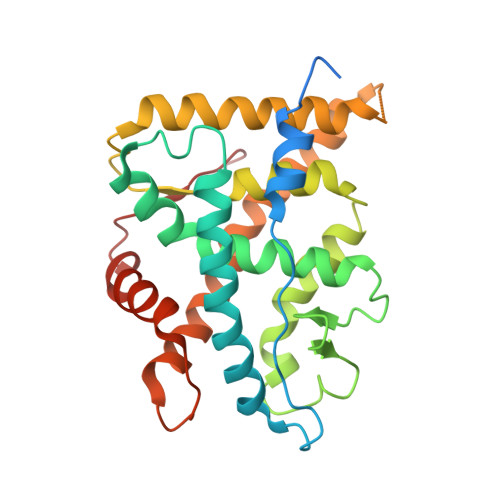
The Three-dimensional Structures of Antagonistic and Agonistic Forms of the Glucocorticoid Receptor Ligand-binding Domain: RU-486 INDUCES A TRANSCONFORMATION THAT LEADS TO ACTIVE ANTAGONISM.
Kauppi, B., Jakob, C., Farnegardh, M., Yang, J., Ahola, H., Alarcon, M., Calles, K., Engstrom, O., Harlan, J., Muchmore, S., Ramqvist, A.-K., Thorell, S., Ohman, L., Greer, J., Gustafsson, J.-A., Carlstedt-Duke, J., Carlquist, M.(2003) J Biological Chem 278: 22748-22754
- PubMed: 12686538 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M212711200
- Primary Citation Related Structures:
1NHZ, 1P93 - PubMed Abstract:
Here we describe the three-dimensional crystal structures of human glucocorticoid receptor ligand-binding domain (GR-LBD) in complex with the antagonist RU-486 at 2.3 A resolution and with the agonist dexamethasone ligand together with a coactivator peptide at 2.8 A. The RU-486 structure was solved in several different crystal forms, two with helix 12 intact (GR1 and GR3) and one with a protease-digested C terminus (GR2). In GR1, part of helix 12 is in a position that covers the co-activator pocket, whereas in the GR3, domain swapping is seen between the crystallographically identical subunits in the GR dimer. An arm consisting of the end of helix 11 and beyond stretches out from one molecule, and helix 12 binds to the other LBD, partly blocking the coactivator pocket of that molecule. This type of GR-LBD dimer has not been described before but might be an artifact from crystallization. Furthermore, the subunits of the GR3 dimers are covalently connected via a disulfide bond between the Cys-736 residues in the two molecules. All three RU-486 GR-LBD structures show that GR has a very flexible region between the end of helix 11 and the end of helix 12.
- Structure Biology, Karo Bio AB, Novum, SE-141 57 Huddinge, Sweden. kauppi@karobio.se
Organizational Affiliation: