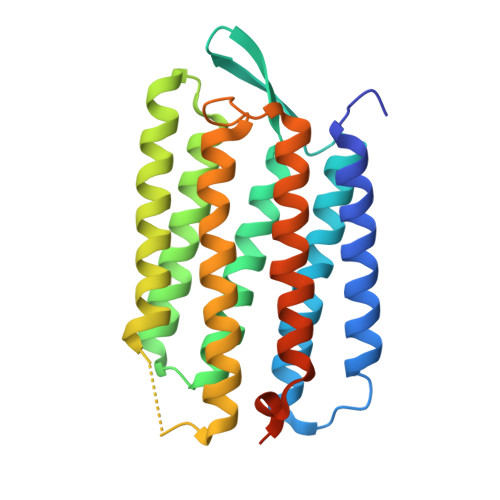
Crystallographic Structures of the M and N Intermediates of Bacteriorhodopsin: Assembly of a Hydrogen-Bonded Chain of Water Molecules between Asp-96 and the Retinal Schiff Base
Schobert, B., Brown, L.S., Lanyi, J.K.(2003) J Mol Biology 330: 553-570
- PubMed: 12842471 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00576-x
- Primary Citation Related Structures:
1P8H, 1P8I, 1P8U - PubMed Abstract:
An M intermediate of wild-type bacteriorhodopsin and an N intermediate of the V49A mutant were accumulated in photostationary states at pH 5.6 and 295 K, and their crystal structures determined to 1.52A and 1.62A resolution, respectively. They appear to be M(1) and N' in the sequence, M(1)<-->M(2)<-->M'(2)<-->N<-->N'-->O-->BR, where M(1), M(2), and M'(2) contain an unprotonated retinal Schiff base before and after a reorientation switch and after proton release to the extracellular surface, while N and N' contain a reprotonated Schiff base, before and after reprotonation of Asp96 from the cytoplasmic surface. In M(1), we detect a cluster of three hydrogen-bonded water molecules at Asp96, not present in the BR state. In M(2), whose structure we reported earlier, one of these water molecules intercalates between Asp96 and Thr46. In N', the cluster is transformed into a single-file hydrogen-bonded chain of four water molecules that connects Asp96 to the Schiff base. We find a network of three water molecules near residue 219 in the crystal structure of the non-illuminated F219L mutant, where the residue replacement creates a cavity. This suggests that the hydration of the cytoplasmic region we observe in N' might have occurred spontaneously, beginning at an existing water molecule as nucleus, in the cavities from residue rearrangements in the photocycle.
- Department of Physiology and Biophysics, University of California, D345 Medical Science I, Irvine, CA 92697, USA.
Organizational Affiliation: