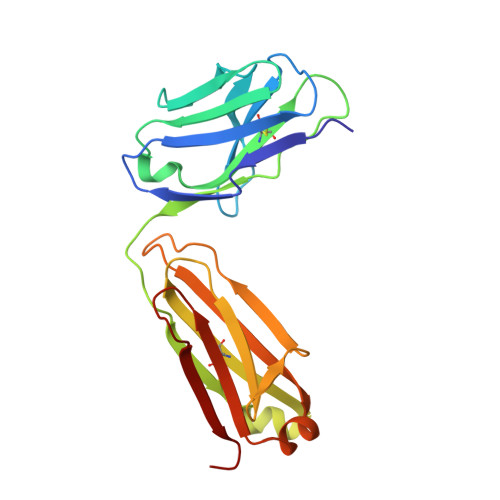
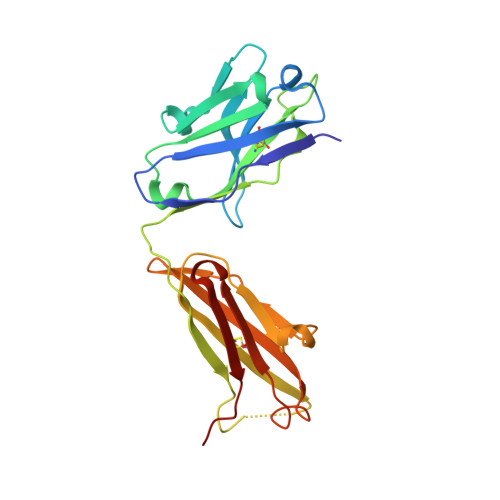
Structure of an anti-DNA fab complexed with a non-DNA ligand provides insights into cross-reactivity and molecular mimicry.
Schuermann, J.P., Henzl, M.T., Deutscher, S.L., Tanner, J.J.(2004) Proteins 57: 269-278
- PubMed: 15340914 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20200
- Primary Citation Related Structures:
1P7K - PubMed Abstract:
Antibodies that recognize DNA (anti-DNA) are part of the autoimmune response underlying systemic lupus erythematosus. To better understand molecular recognition by anti-DNA antibodies, crystallographic studies have been performed using an anti-ssDNA antigen-binding fragment (Fab) known as DNA-1. The previously determined structure of a DNA-1/dT5 complex revealed that thymine bases insert into a narrow groove, and that ligand recognition primarily involves the bases of DNA. We now report the 1.75-A resolution structure of DNA-1 complexed with the biological buffer HEPES (4-(2-Hydroxyethyl)piperazine-1-ethanesulfonic acid). All three light chain complementarity-determining regions (CDRs) and HCDR3 contribute to binding. The HEPES sulfonate hydrogen bonds to His L91, Asn L50, and to the backbone of Tyr H100 and Tyr H100A. The Tyr side-chains of L32, L92, H100, and H100A form nonpolar contacts with the HEPES ethylene and piperazine groups. Comparison to the DNA-1/dT5 structure reveals that the dual recognition of dT5 and HEPES requires a 13-A movement of HCDR3. This dramatic structural change converts the combining site from a narrow groove, appropriate for the edge-on insertion of thymine bases, to one sufficiently wide to accommodate the HEPES sulfonate and piperazine. Isothermal titration calorimetry verified the association of HEPES with DNA-1 under conditions similar those used for crystallization (2 M ammonium sulfate). Interestingly, the presence of 2 M ammonium sulfate increases the affinities of DNA-1 for both HEPES and dT5, suggesting that non-polar Fab-ligand interactions are important for molecular recognition in highly ionic solvent conditions. The structural and thermodynamic data suggest a molecular mimicry mechanism based on structural plasticity and hydrophobic interactions.
- Department of Chemistry, University of Missouri-Columbia, Columbia, Missouri 65211, USA.
Organizational Affiliation: