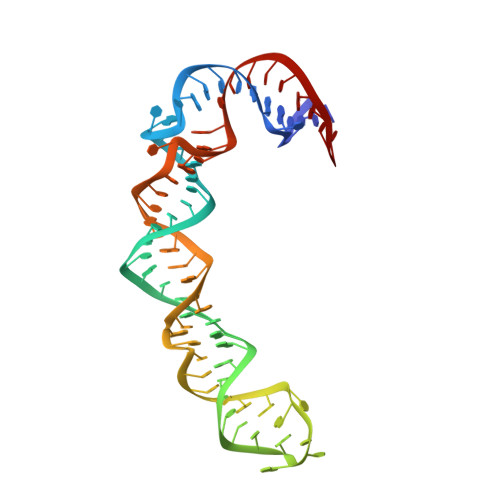
Structure of HCV IRES domain II determined by NMR.
Lukavsky, P.J., Kim, I., Otto, G.A., Puglisi, J.D.(2003) Nat Struct Biol 10: 1033-1038
- PubMed: 14578934 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1004
- Primary Citation Related Structures:
1P5M, 1P5N, 1P5O, 1P5P - PubMed Abstract:
Complex RNA structures regulate many biological processes, but are often too large for structure determination by NMR methods. The 5' untranslated region (5' UTR) of the hepatitis C viral (HCV) RNA genome contains an internal ribosome entry site (IRES) that binds to 40S ribosomal subunits with high affinity and specificity to control translation. Domain II of the HCV IRES forms a 25-kDa folded subdomain that may alter ribosome conformation. We report here the structure of domain II as determined using an NMR approach that combines short- and long-range structural data. Domain II adopts a distorted L-shape structure, and its overall shape in the free form is markedly similar to its 40S subunit-bound form; this suggests how domain II may modulate 40S subunit conformation. The results show how NMR can be used for structural analysis of large biological RNAs.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, California 94305-5126, USA.
Organizational Affiliation: