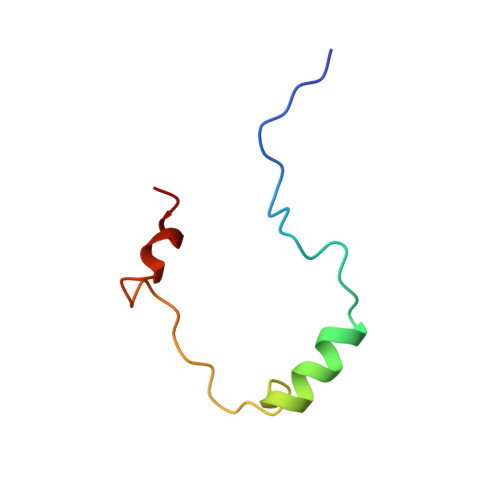
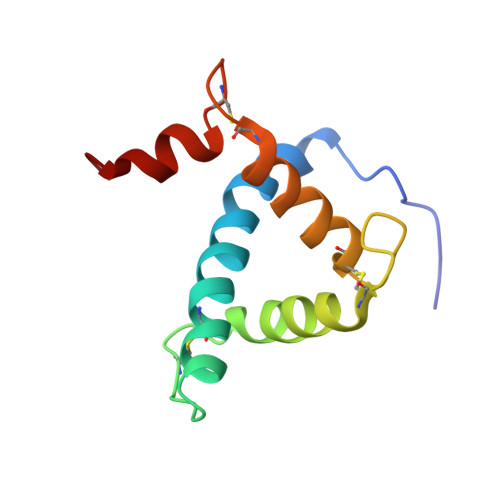
Structural basis for negative regulation of hypoxia-inducible factor-1alpha by CITED2.
Freedman, S.J., Sun, Z.Y., Kung, A.L., France, D.S., Wagner, G., Eck, M.J.(2003) Nat Struct Biol 10: 504-512
- PubMed: 12778114 Search on PubMed
- DOI: https://doi.org/10.1038/nsb936
- Primary Citation Related Structures:
1P4Q - PubMed Abstract:
Expression of hypoxia-responsive genes is mediated by the heterodimeric transcription factor hypoxia-inducible factor-1 (HIF-1) in complex with the p300/CREB-binding protein (p300/CBP) transcriptional coactivator. The protein CITED2, which binds p300/CBP, is thought to be a negative regulator of HIF-1 transactivation. We show that the CITED2 transactivation domain (TAD) disrupts a complex of the HIF-1alpha C-terminal TAD (C-TAD) and the cysteine-histidine-rich 1 (CH1) domain of p300/CBP by binding CH1 with high affinity. The high-resolution solution structure of the CITED2 TAD-p300 CH1 complex shows that the CITED2 TAD, like the HIF-1alpha C-TAD, folds on a helical, Zn2+-containing CH1 scaffold. The CITED2 TAD binds a different, more extensive surface of CH1 than does the HIF-1alpha C-TAD. However, a conserved 'LPXL' sequence motif in CITED2 and HIF-1alpha interacts with an overlapping binding site on CH1. Mutation of the LPEL sequence in full-length CITED2 abolishes p300 binding in vivo. These findings reveal that CITED2 regulates HIF-1 by competing for a hot spot on the p300 CH1 domain.
- Division of Hemostasis and Thrombosis, Beth Israel Deaconess Medical Center, 41 Avenue Louis Pasteur, Boston, Massachusetts 02115, USA.
Organizational Affiliation: