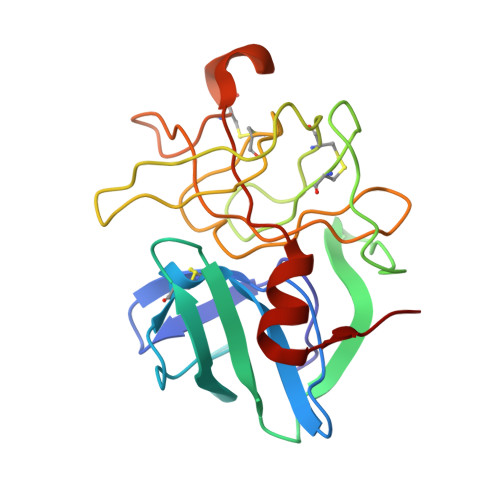
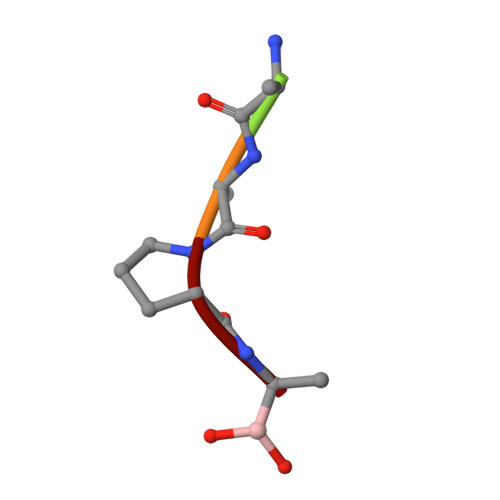
Structural analysis of specificity: alpha-lytic protease complexes with analogues of reaction intermediates.
Bone, R., Frank, D., Kettner, C.A., Agard, D.A.(1989) Biochemistry 28: 7600-7609
- PubMed: 2611204 Search on PubMed
- DOI: https://doi.org/10.1021/bi00445a015
- Primary Citation Related Structures:
1P02, 1P03, 1P04, 1P05, 1P06 - PubMed Abstract:
To better understand the structural basis of enzyme specificity, the structures of complexes formed between alpha-lytic protease, an extracellular serine protease of Lysobacter enzymogenes, and five inhibitory peptide boronic acids (R2-boroX, where R2 is methoxysuccinyl-Ala-Ala-Pro- and boroX is the alpha-aminoboronic acid analogue of Ala, Val, Ile, Norleu, or Phe) have been studied at high resolution by X-ray crystallography. The enzyme has primary specificity for Ala in the P1 position of peptide substrates with catalytic efficiency decreasing with increasing side-chain volume. Enzyme affinity for inhibitors with boroVal, boroIle, and boroPhe residues is much higher than expected on the basis of the catalytic efficiencies of homologous substrates. Covalent tetrahedral adducts are formed between the active-site serine and the boronic acid moieties of R2-boroAla, R2-boroVal R2-boroIle, and R2-boroNorleu. Though R2-boroVal is a slowly bound inhibitor and R2-boroAla is rapidly bound [Kettner, C. A., Bone, R., Agard, D. A., & Bachovchin, W. W. (1988) Biochemistry 27, 7682-7688], there appear to be no structural differences that could account for slow binding. The removal from solution of 20% more hydrophobic surface on binding accounts for the improved affinity of alpha-lytic protease for R2-boroVal relative to R2-boroAla. The high affinity of the enzyme for R2-boroIle derives from the selective binding of the L-allo stereoisomer of the boroIle residue, which can avoid bad steric interactions in the binding pocket.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Biochemistry and Biophysics, University of California, San Francisco 94143-0448.
Organizational Affiliation: