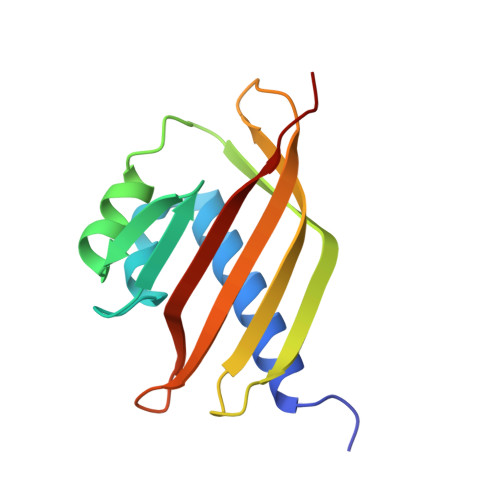
The 1.6 angstroms resolution crystal structure of nuclear transport factor 2 (NTF2).
Bullock, T.L., Clarkson, W.D., Kent, H.M., Stewart, M.(1996) J Mol Biology 260: 422-431
- PubMed: 8757804 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0411
- Primary Citation Related Structures:
1OUN - PubMed Abstract:
Nuclear transport factor 2 (NTF2) facilitates protein transport into the nucleus and interacts with both the small Ras-like GTPase Ran and nucleoporin p62. We have determined the structure of bacterially expressed rat NTF2 at 1.6 angstroms resolution using X-ray crystallography. The NTF2 polypeptide chain forms an alpha + beta barrel that opens at one end to form a distinctive hydrophobic cavity and its fold is homologous to that of scytalone dehydratase. The NTF2 hydrophobic cavity is a candidate for a potential binding site for other proteins involved in nuclear import such as Ran and nucleoporin p62. In addition, the hydrophobic cavity contains a putative catalytic Asp-His pair, which raises the possibility of an unanticipated enzymatic activity of the molecule that may have implications for the molecular mechanism of nuclear protein import.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: