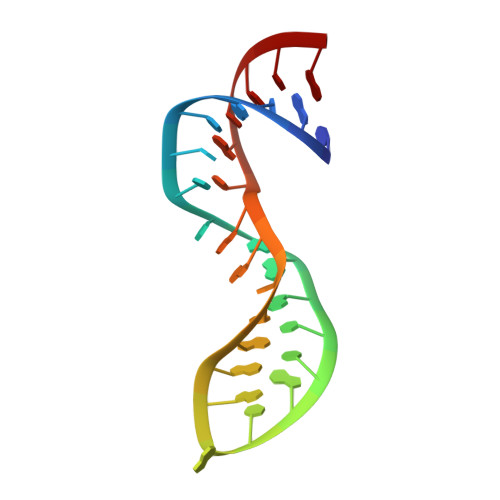
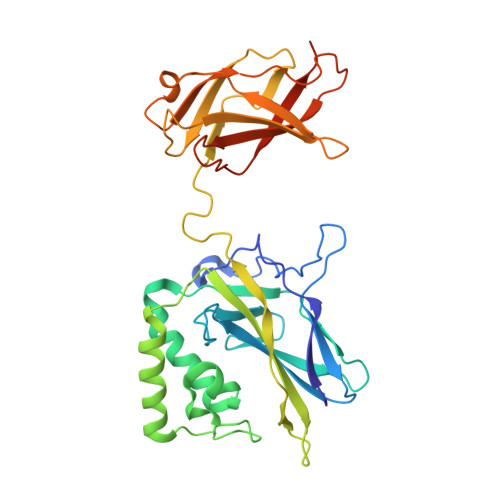
Crystal structure of NF-kappaB (p50)2 complexed to a high-affinity RNA aptamer.
Huang, D.B., Vu, D., Cassiday, L.A., Zimmerman, J.M., Maher III, L.J., Ghosh, G.(2003) Proc Natl Acad Sci U S A 100: 9268-9273
- PubMed: 12886018 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1632011100
- Primary Citation Related Structures:
1OOA - PubMed Abstract:
We have recently identified an RNA aptamer for the transcription factor NF-kappaB p50 homodimer [(p50)2], which exhibits little sequence resemblance to the consensus DNA target for (p50)2, but binds (p50)2 with an affinity similar to that of the optimal DNA target. We describe here the 2.45-A resolution x-ray crystal structure of the p50 RHR/RNA aptamer complex. The structure reveals that two RNA molecules bind independent of each other to the p50 N-terminal Ig-like domains. The RNA secondary structure is comprised of a stem and a stem-loop separated by an internal loop folded into a kinked helix because of the cross-strand stacking of three internal loop guanines. These guanines, placed at the edge of the 3' helix, together with the major groove of the irregular 3' helix, form the binding surface for p50. Each p50 monomer uses the same surface to recognize the distorted RNA major groove as observed in the kappaB DNA/p50 RHR complex, resulting in strikingly similar interfaces. The structure reveals how the aptamer specifically selects p50 and discriminates against p65. We also discuss the physiological implications of RNA binding by (p50)2.
- Department of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 93092-0359, USA.
Organizational Affiliation: