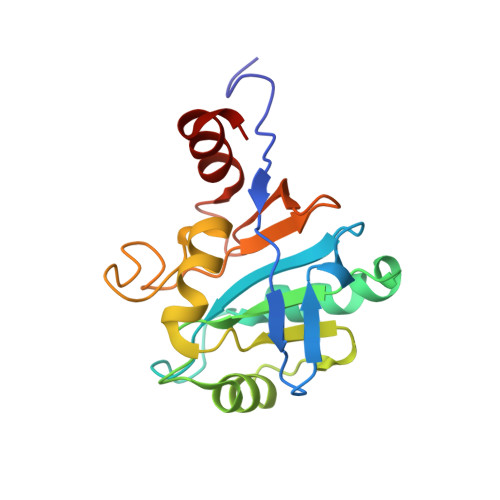
Solution Structure of Sco1: A Thioredoxin-like Protein Involved in Cytochrome c Oxidase Assembly
Balatri, E., Banci, L., Bertini, I., Cantini, F., Ciofi-Baffoni, S.(2003) Structure 11: 1431-1433
- PubMed: 14604533 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2003.10.004
- Primary Citation Related Structures:
1ON4 - PubMed Abstract:
Sco1, a protein required for the proper assembly of cytochrome c oxidase, has a soluble domain anchored to the cytoplasmic membrane through a single transmembrane segment. The solution structure of the soluble part of apoSco1 from Bacillus subtilis has been solved by NMR and the internal mobility characterized. Its fold places Sco1 in a distinct subgroup of the functionally unrelated thioredoxin proteins. In vitro Sco1 binds copper(I) through a CXXXCP motif and possibly His 135 and copper(II) in two different species, thus suggesting that copper(II) is adventitious more than physiological. The Sco1 structure represents the first structure of this class of proteins, present in a variety of eukaryotic and bacterial organisms, and elucidates a link between copper trafficking proteins and thioredoxins. The availability of the structure has allowed us to model the homologs Sco1 and Sco2 from S. cerevisiae and to discuss the physiological role of the Sco family.
- Magnetic Resonance Center CERM and Department of Chemistry, University of Florence, Via Luigi Sacconi 6, 50019 Sesto Fiorentino, Florence, Italy.
Organizational Affiliation: