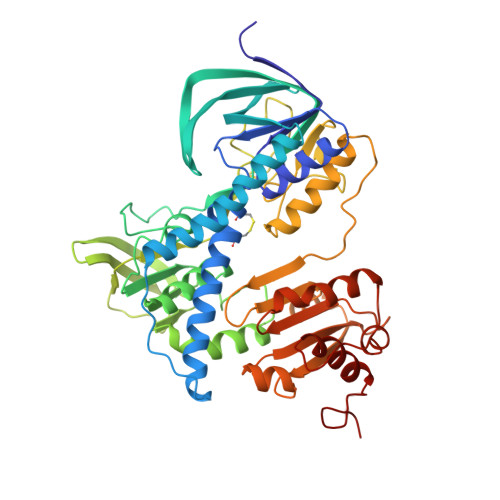
Molecular structure of the lipoamide dehydrogenase domain of a surface antigen from Neisseria meningitidis.
Li de la Sierra, I., Pernot, L., Prange, T., Saludjian, P., Schiltz, M., Fourme, R., Padron, G.(1997) J Mol Biology 269: 129-141
- PubMed: 9193005 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1009
- Primary Citation Related Structures:
1OJT - PubMed Abstract:
The protein p64k from the surface of the Neisseria meningitidis bacteria has been characterized as a two-domain protein. It contains a dihydrolipoamide dehydrogenase domain of 482 residues, involving a FAD prosthetic group as a cofactor, and a smaller lipoic acid binding domain of 86 residues. The two domains are joined by a flexible segment rich in alanine and proline residues. The structure of the dihydrolipoamide dehydrogenase domain was determined by X-ray diffraction. It was solved by a combination of molecular replacement and multiple isomorphous replacement techniques and refined to 2.7 A resolution. In the crystal, the recombinant p64k mimics the functional homo-dimer by using one of the crystallographic 2-fold axes. The reactive disulphide bridge Cys161-Cys166 is in the oxidised state and the FAD is bound in an extended conformation. This main domain contains the major antigenic determinant of the protein, an extended loop of 32 residues at the surface of the protein. A mis-attribution at residue 553 in the sequence has been detected by inspection of electron density maps and the geometry. However, when compared to the other dihydrolipoamide dehydrogenases, there are some significant differences: (1) an unusual number of cis-proline residues and (2) a new motif built around a 2-fold axis by the sulphur atoms of residues Met558, Cys560 and their symmetry related equivalents.
- CIGB, Cubanacan, La Habana, Cuba.
Organizational Affiliation: