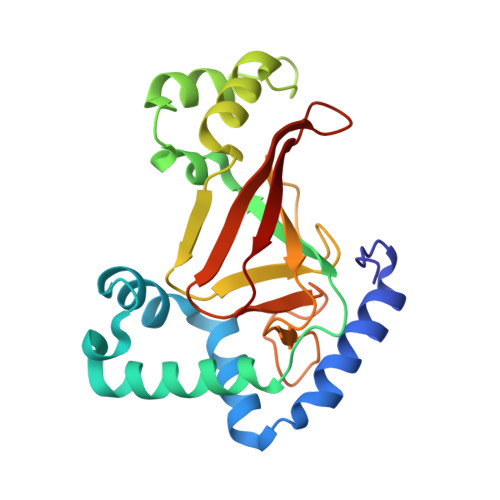
The Crystal Structure of C3Stau2 from Staphylococcus Aureus and its Complex with Nad
Evans, H.R., Sutton, J.M., Holloway, D.E., Ayriss, J., Shone, C.C., Acharya, K.R.(2003) J Biological Chem 278: 45924
- PubMed: 12933793
- DOI: https://doi.org/10.1074/jbc.M307719200
- Primary Citation of Related Structures:
1OJQ, 1OJZ - PubMed Abstract:
The C3stau2 exoenzyme from Staphylococcus aureus is a C3-like ADP-ribosyltransferase that ADP-ribosylates not only RhoA-C but also RhoE/Rnd3. In this study we have crystallized and determined the structure of C3stau2 in both its native form and in complex with NAD at 1.68- and 2.02-A resolutions, respectively. The topology of C3stau2 is similar to that of C3bot1 from Clostridium botulinum (with which it shares 35% amino acid sequence identity) with the addition of two extra helices after strand beta1. The native structure also features a novel orientation of the catalytic ARTT loop, which approximates the conformation seen for the "NAD bound" form of C3bot1. C3stau2 orients NAD similarly to C3bot1, and on binding NAD, C3stau2 undergoes a clasping motion and a rearrangement of the phosphate-nicotinamide binding loop, enclosing the NAD in the binding site. Comparison of these structures with those of C3bot1 and related toxins reveals a degree of divergence in the interactions with the adenine moiety among the ADP-ribosylating toxins that contrasts with the more conserved interactions with the nicotinamide. Comparison with C3bot1 gives some insight into the different protein substrate specificities of these enzymes.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: