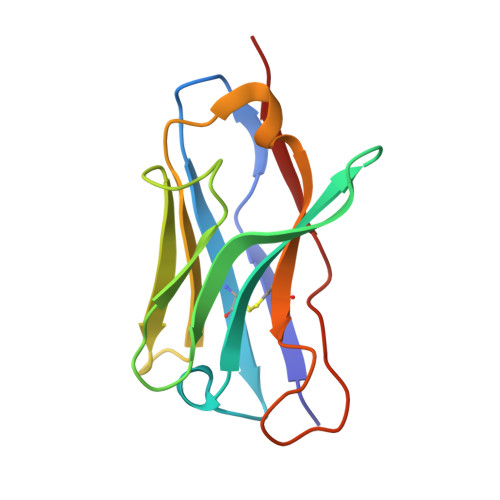
Crystal Structure of Hel4, a Soluble, Refoldable Human V(H) Single Domain with a Germ-Line Scaffold
Jespers, L., Schon, O., James, L.C., Veprintsev, D., Winter, G.(2004) J Mol Biology 337: 893
- PubMed: 15033359
- DOI: https://doi.org/10.1016/j.jmb.2004.02.013
- Primary Citation Related Structures:
1OHQ - PubMed Abstract:
The antigen binding site of antibodies usually comprises associated heavy (V(H)) and light (V(L)) chain variable domains, but in camels and llamas, the binding site frequently comprises the heavy chain variable domain only (referred to as V(HH)). In contrast to reported human V(H) domains, V(HH) domains are well expressed from bacteria and yeast, are readily purified in soluble form and refold reversibly after heat-denaturation. These desirable properties have been attributed to highly conserved substitutions of the hydrophobic residues of V(H) domains, which normally interact with complementary V(L) domains. Here, we describe the discovery and characterisation of an isolated human V(H) domain (HEL4) with properties similar to those of V(HH) domains. HEL4 is highly soluble at concentrations of > or =3 mM, essentially monomeric and resistant to aggregation upon thermodenaturation at concentrations as high as 56 microM. However, in contrast to V(HH) domains, the hydrophobic framework residues of the V(H):V(L) interface are maintained and the only sequence changes from the corresponding human germ-line segment (V3-23/DP-47) are located in the loops comprising the complementarity determining regions (CDRs). The crystallographic structure of HEL4 reveals an unusual feature; the side-chain of a framework residue (Trp47) is flipped into a cavity formed by Gly35 of CDR1, thereby increasing the hydrophilicity of the V(H):V(L) interface. To evaluate the specific contribution of Gly35 to domain properties, Gly35 was introduced into a V(H) domain with poor solution properties. This greatly enhanced the recovery of the mutant from a gel filtration matrix, but had little effect on its ability to refold reversibly after heat denaturation. Our results confirm the importance of a hydrophilic V(H):V(L) interface for purification of isolated V(H) domains, and constitute a step towards the design of isolated human V(H) domains with practical properties for immunotherapy.
- Laboratory of Molecular Biology, Medical Research Council Centre, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: