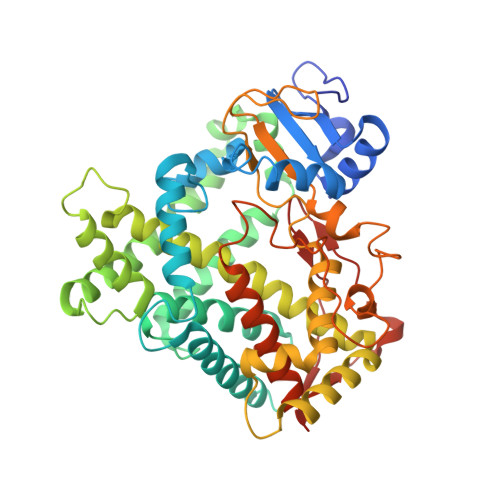
Crystal Structure of Human Cytochrome P450 2C9 with Bound Warfarin
Williams, P.A., Cosme, J., Ward, A., Angove, H.C., Matak Vinkovic, D., Jhoti, H.(2003) Nature 424: 464
- PubMed: 12861225
- DOI: https://doi.org/10.1038/nature01862
- Primary Citation of Related Structures:
1OG2, 1OG5 - PubMed Abstract:
Cytochrome P450 proteins (CYP450s) are membrane-associated haem proteins that metabolize physiologically important compounds in many species of microorganisms, plants and animals. Mammalian CYP450s recognize and metabolize diverse xenobiotics such as drug molecules, environmental compounds and pollutants. Human CYP450 proteins CYP1A2, CYP2C9, CYP2C19, CYP2D6 and CYP3A4 are the major drug-metabolizing isoforms, and contribute to the oxidative metabolism of more than 90% of the drugs in current clinical use. Polymorphic variants have also been reported for some CYP450 isoforms, which has implications for the efficacy of drugs in individuals, and for the co-administration of drugs. The molecular basis of drug recognition by human CYP450s, however, has remained elusive. Here we describe the crystal structure of a human CYP450, CYP2C9, both unliganded and in complex with the anti-coagulant drug warfarin. The structure defines unanticipated interactions between CYP2C9 and warfarin, and reveals a new binding pocket. The binding mode of warfarin suggests that CYP2C9 may undergo an allosteric mechanism during its function. The newly discovered binding pocket also suggests that CYP2C9 may simultaneously accommodate multiple ligands during its biological function, and provides a possible molecular basis for understanding complex drug-drug interactions.
- Astex Technology, 436 Cambridge Science Park, Milton Road, Cambridge CB4 0QA, UK.
Organizational Affiliation: