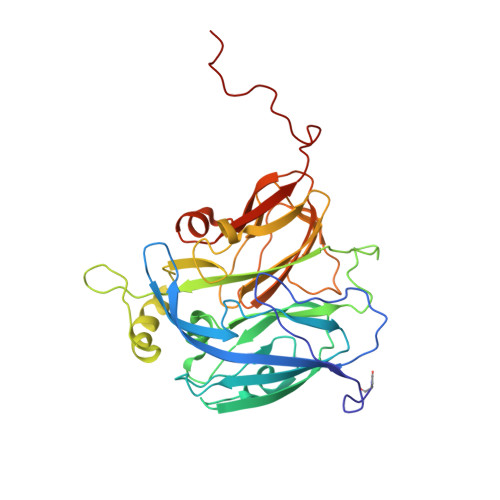
Atomic Resolution Structures of Native Copper Nitrite Reductase from Alcaligenes Xylosoxidans and the Active Site Mutant Asp92Glu
Ellis, M.J., Dodd, F.E., Sawers, G., Eady, R.R., Hasnain, S.S.(2003) J Mol Biology 328: 429
- PubMed: 12691751 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00308-5
- Primary Citation Related Structures:
1OE1, 1OE2, 1OE3 - PubMed Abstract:
We provide the first atomic resolution (<1.20 A) structure of a copper protein, nitrite reductase, and of a mutant of the catalytically important Asp92 residue (D92E). The atomic resolution where carbon-carbon bonds of the peptide become clearly resolved, remains a key goal of structural analysis. Despite much effort and technological progress, still very few structures are known at such resolution. For example, in the Protein Data Bank (PDB) there are some 200 structures of copper proteins but the highest resolution structure is that of amicyanin, a small (12 kDa) protein, which has been resolved to 1.30 A. Here, we present the structures of wild-type copper nitrite reductase (wtNiR) from Alcaligenes xylosoxidans (36.5 kDa monomer), the "half-apo" recombinant native protein and the D92E mutant at 1.04, 1.15 and 1.12A resolutions, respectively. These structures provide the basis from which to build a detailed mechanism of this important enzyme.
- Faculty of Applied Science, De Montfort University, Leicester LE1 9BH, UK.
Organizational Affiliation: