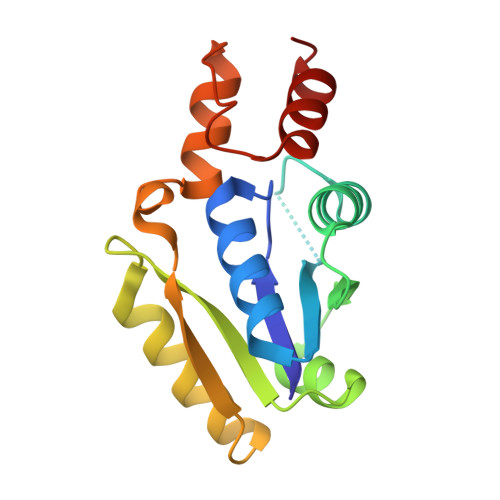
Structure and Implications for the Thermal Stability of Phosphopantetheine Adenylyltransferase from Thermus Thermophilus.
Takahashi, H., Inagaki, E., Fujimoto, Y., Kuroishi, C., Nodake, Y., Nakamura, Y., Arisaka, F., Yutani, K., Kuramitsu, S., Yokoyama, S., Yamamoto, M., Miyano, M., Tahirov, T.H.(2004) Acta Crystallogr D Biol Crystallogr 60: 97
- PubMed: 14684898 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444903025319
- Primary Citation Related Structures:
1OD6 - PubMed Abstract:
Phosphopantetheine adenylyltransferase (PPAT) is an essential enzyme in bacteria that catalyzes the rate-limiting step in coenzyme A (CoA) biosynthesis by transferring an adenylyl group from ATP to 4'-phosphopantetheine (Ppant), yielding 3'-dephospho-CoA (dPCoA). The crystal structure of PPAT from Thermus thermophilus HB8 (Tt PPAT) complexed with Ppant has been determined by the molecular-replacement method at 1.5 A resolution. The overall fold of the enzyme is almost the same as that of Escherichia coli PPAT, a hexamer having point group 32. The asymmetric unit of Tt PPAT contains a monomer and the crystallographic triad and dyad coincide with the threefold and twofold axes of the hexamer, respectively. Most of the important atoms surrounding the active site in E. coli PPAT are conserved in Tt PPAT, indicating similarities in their substrate binding and enzymatic reaction. The notable difference between E. coli PPAT and Tt PPAT is the simultaneous substrate recognition by all six subunits of Tt PPAT compared with substrate recognition by only three subunits in E. coli PPAT. Comparative analysis also revealed that the higher stability of Tt PPAT arises from stabilization of each subunit by hydrophobic effects, hydrogen bonds and entropic effects.
- Highthroughput Factory, RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: