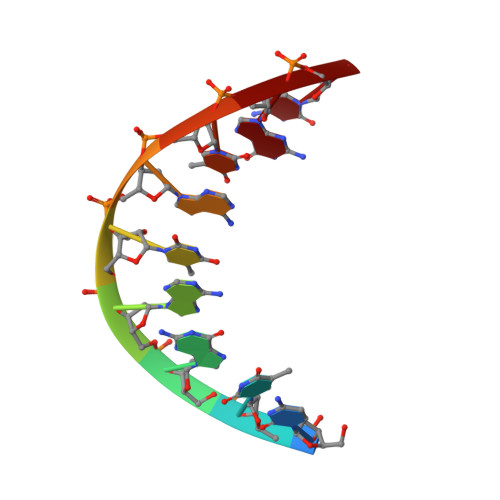
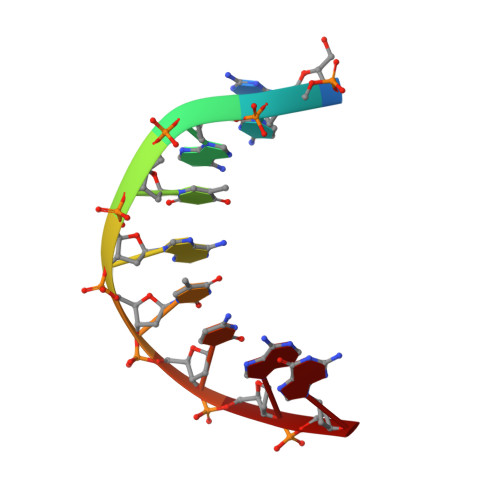
NMR Solution Structure of a DsDNA Containing a Bicyclic D-Arabino-Configured Nucleotide Fixed in an O4'-Endo Sugar Conformation
Tommerholt, H.V., Christensen, N.K., Nielsen, P., Wengel, J., Stein, P.C., Jacobsen, J.P., Petersen, M.(2003) Org Biomol Chem 1: 1790-1797
- PubMed: 12926371
- DOI: https://doi.org/10.1039/b300848g
- Primary Citation Related Structures:
1OCI - PubMed Abstract:
[3.2.0]bcANA is a D-arabino-configured bicyclic nucleotide with a 2'-O,3'-C-methylene bridge. We here present the high-resolution NMR structure of a [3.2.0]bcANA modified dsDNA nonamer with one modified nucleotide incorporated. NOE restraints were obtained by analysis of NOESY cross peak intensities using a full relaxation matrix approach, and subsequently these restraints were incorporated into a simulated annealing scheme for the structure determination. In addition, the furanose ring puckers of the deoxyribose moieties were determined by analysis of COSY cross peaks. The modified duplex adopts a B-like geometry with Watson-Crick base pairing in all base pairs and all glycosidic angles in the anti range. The stacking arrangement of the nucleobases appears to be unperturbed relative to the normal B-like arrangement. The 2'-O,3'-C-methylene bridge of the modified nucleotide is located at the brim of the major groove where it fits well into the B-type duplex framework. The sugar pucker of the [3.2.0]bcANA nucleotide is O4'-endo and this sugar conformation causes a change in the delta backbone angle relative to the C2'-endo deoxyribose sugar pucker. This change is absorbed locally by slight changes in the epsilon and zeta angles of the modified nucleotide. Overall, the [3.2.0]bcANA modifications fits very well into a B-like duplex framework and only small and local perturbations are observed relative to the unmodified dsDNA of identical base sequence.
- Department of Chemistry, University of Southern Denmark, DK-5230 Odense M, Denmark.
Organizational Affiliation: