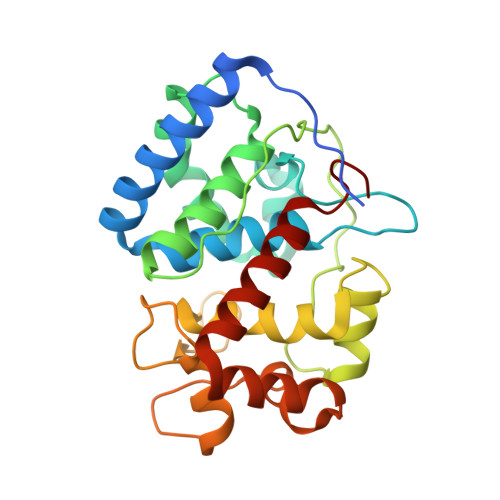
Crystal Structure of the Ascorbate Peroxidase-Ascorbate Complex
Sharp, K.H., Mewies, M., Moody, P.C.E., Raven, E.L.(2003) Nat Struct Biol 10: 303
- PubMed: 12640445 Search on PubMed
- DOI: https://doi.org/10.1038/nsb913
- Primary Citation Related Structures:
1OAF, 1OAG - PubMed Abstract:
Heme peroxidases catalyze the H2O2-dependent oxidation of a variety of substrates, most of which are organic. Mechanistically, these enzymes are well characterized: they share a common catalytic cycle that involves formation of a two-electron, oxidized Compound I intermediate followed by two single-electron reduction steps by substrate. The substrate specificity is more diverse--most peroxidases oxidize small organic substrates, but there are prominent exceptions--and there is a notable absence of structural information for a representative peroxidase-substrate complex. Thus, the features that control substrate specificity remain undefined. We present the structure of the complex of ascorbate peroxidase-ascorbate. The structure defines the ascorbate-binding interaction for the first time and provides new rationalization of the unusual functional features of the related cytochrome c peroxidase enzyme, which has been a benchmark for peroxidase catalysis for more than 20 years. A new mechanism for electron transfer is proposed that challenges existing views of substrate oxidation in other peroxidases.
- Department of Chemistry, University of Leicester, University Road, Leicester, LE1 7RH, England, UK.
Organizational Affiliation: