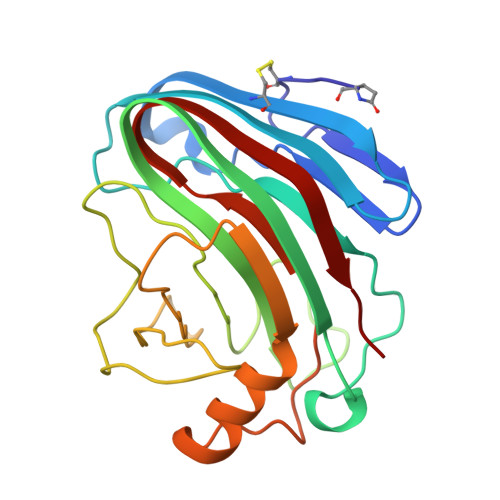
Comparison of Family 12 Glycoside Hydrolases and Recruited Substitutions Important for Thermal Stability
Sandgren, M., Gualfetti, P.J., Shaw, A., Gross, L.S., Saldajeno, M., Day, A.G., Jones, T.A., Mitchinson, C.(2003) Protein Sci 12: 848
- PubMed: 12649442 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.0237703
- Primary Citation Related Structures:
1OA2, 1OA3, 1OA4 - PubMed Abstract:
As part of a program to discover improved glycoside hydrolase family 12 (GH 12) endoglucanases, we have studied the biochemical diversity of several GH 12 homologs. The H. schweinitzii Cel12A enzyme differs from the T. reesei Cel12A enzyme by only 14 amino acids (93% sequence identity), but is much less thermally stable. The bacterial Cel12A enzyme from S. sp. 11AG8 shares only 28% sequence identity to the T. reesei enzyme, and is much more thermally stable. Each of the 14 sequence differences from H. schweinitzii Cel12A were introduced in T. reesei Cel12A to determine the effect of these amino acid substitutions on enzyme stability. Several of the T. reesei Cel12A variants were found to have increased stability, and the differences in apparent midpoint of thermal denaturation (T(m)) ranged from a 2.5 degrees C increase to a 4.0 degrees C decrease. The least stable recruitment from H. schweinitzii Cel12A was A35S. Consequently, the A35V substitution was recruited from the more stable S. sp. 11AG8 Cel12A and this T. reesei Cel12A variant was found to have a T(m) 7.7 degrees C higher than wild type. Thus, the buried residue at position 35 was shown to be of critical importance for thermal stability in this structural family. There was a ninefold range in the specific activities of the Cel12 homologs on o-NPC. The most and least stable T. reesei Cel12A variants, A35V and A35S, respectively, were fully active. Because of their thermal tolerance, S. sp. 11AG8 Cel12A and T. reesei Cel12A variant A35V showed a continual increase in activity over the temperature range of 25 degrees C to 60 degrees C, whereas the less stable enzymes T. reesei Cel12A wild type and the destabilized A35S variant, and H. schweinitzii Cel12A showed a decrease in activity at the highest temperatures. The crystal structures of the H. schweinitzii, S. sp. 11AG8, and T. reesei A35V Cel12A enzymes have been determined and compared with the wild-type T. reesei Cel12A enzyme. All of the structures have similar Calpha traces, but provide detailed insight into the nature of the stability differences. These results are an example of the power of homolog recruitment as a method for identifying residues important for stability.
- Department of Cell and Molecular Biology, Uppsala University, Biomedical Center, SE-75124 Uppsala, Sweden.
Organizational Affiliation: