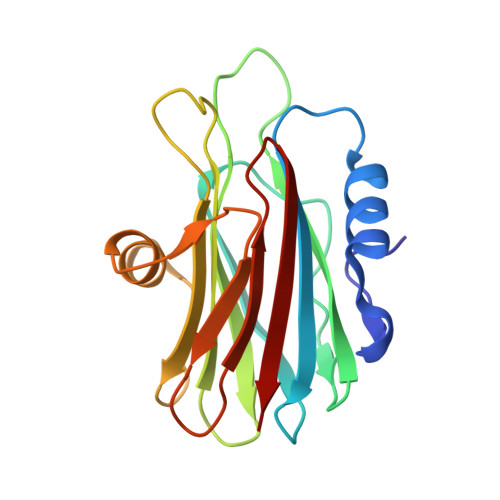
Crystal and Electron Microscopy Structures of Sticholysin II Actinoporin Reveal Insights Into the Mechanism of Membrane Pore Formation
Mancheno, J.M., Martin-Benito, J., Martinez-Ripoll, M., Gavilanes, J.G., Hermoso, J.A.(2003) Structure 11: 1319
- PubMed: 14604522 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2003.09.019
- Primary Citation Related Structures:
1GWY, 1O71, 1O72 - PubMed Abstract:
Sticholysin II (StnII) is a pore-forming protein (PFP) produced by the sea anemone Stichodactyla helianthus. We found out that StnII exists in a monomeric soluble state but forms tetramers in the presence of a lipidic interface. Both structures have been independently determined at 1.7 A and 18 A resolution, respectively, by using X-ray crystallography and electron microscopy of two-dimensional crystals. Besides, the structure of soluble StnII complexed with phosphocholine, determined at 2.4 A resolution, reveals a phospholipid headgroup binding site, which is located in a region with an unusually high abundance of aromatic residues. Fitting of the atomic model into the electron microscopy density envelope suggests that while the beta sandwich structure of the protein remains intact upon oligomerization, the N-terminal region and a flexible and highly basic loop undergo significant conformational changes. These results provide the structural basis for the membrane recognition step of actinoporins and unexpected insights into the oligomerization step.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto Rocasolano, CSIC, Serrano 119, 28006 Madrid, Spain.
Organizational Affiliation: