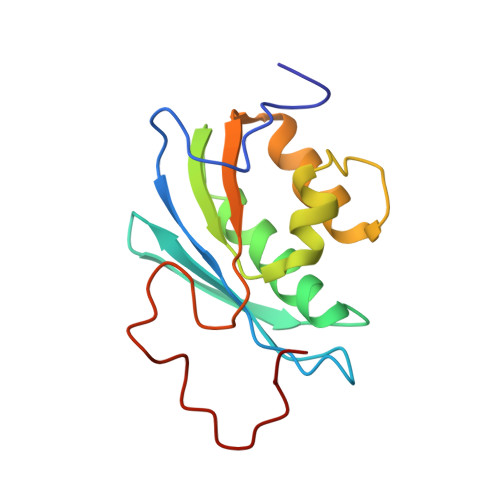
Solution Structure of the Rnase H Domain of the HIV-1 Reverse Transcriptase in the Presence of Magnesium
Pari, K., Mueller, G.A., Derose, E.F., Kirby, T.W., London, R.E.(2003) Biochemistry 42: 639-650
- PubMed: 12534276 Search on PubMed
- DOI: https://doi.org/10.1021/bi0204894
- Primary Citation Related Structures:
1O1W - PubMed Abstract:
This paper presents the first solution structure of the RNase H domain of HIV-1 reverse transcriptase (RT) determined by NMR methods. The solution conditions in this study were at physiological pH in the presence of Mg(2+). An investigation of the dependence of the (1)H-(15)N HSQC spectrum of the RNase H domain on [Mg(2+)] indicates that Mg(2+) produces significant, global effects on the amide chemical shifts, implying that divalent metal ion binding is important for stabilizing the structure of the isolated domain in solution. Analysis of amide shift data as a function of MgCl(2) concentration using either a single- or two-site binding model indicated that the latter provided a significantly improved fit, with the K(D) for site A = 2.7-3.2 mM and K(D) for site B approximately 35 mM, calculated on the assumption that site A is already occupied. Resonances of the [U-(13)C,(15)N]RNase H domain, measured at pH 6.8, in 80 mM MgCl(2), were assigned and NOESY data collected in order to determine the structure. Assignment of the NOESY spectra using the ARIA program resulted in a high-resolution structure for residues 6-114 which was similar to the crystal structure of the isolated domain,. The data were insufficient to define a compact structure for the C-terminal residues after 114. Residues I134-L138 located at the C-terminus are highly disordered and give rise to relatively sharp and intense amide resonances, while the amide resonances for the segment from E124 to A132 appear to be largely absent and are presumably subject to significant exchange broadening between different conformational states. Comparisons with crystal structure data for the full reverse transcriptase molecule indicate that the corresponding region is absent in nearly all of the crystal structures determined for the P2(1)2(1)2(1) space group, while these residues adopt an alpha-helix in structures determined for other symmetry groups. This structural heterogeneity indicates that significant conformational variability exists for this segment of the full reverse transcriptase enzyme as well, and the structure of the C-terminal peptide can be selected or deselected, depending on crystallization conditions. This analysis, along with the structural characterization contained herein, challenges the previous paradigm that the dynamic behavior of the isolated RNase H domain differs substantially from the behavior in the intact enzyme. The poor Mg(2+) binding and conformational flexibility of residues located near the active site indicate that substrate binding is a precondition for metal ion binding and for selecting the active site conformation of the RNase H domain.
- Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, P.O. Box 12233, Research Triangle Park, North Carolina 27709, USA.
Organizational Affiliation: