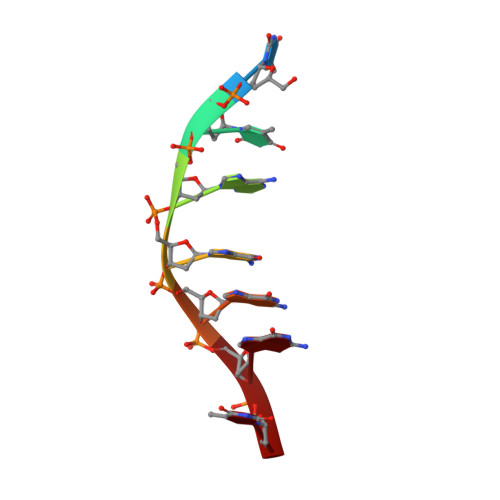
Drug Recognition and Stabilisation of the Parallel-stranded DNA Quadruplex d(TTAGGGT)4 Containing the Human Telomeric Repeat
Gavathiotis, E., Heald, R.A., Stevens, M.F.G., Searle, M.S.(2003) J Mol Biology 334: 25-36
- PubMed: 14596797
- DOI: https://doi.org/10.1016/j.jmb.2003.09.018
- Primary Citation Related Structures:
1NZM - PubMed Abstract:
The NMR structure of the parallel-stranded DNA quadruplex d(TTAGGGT)(4), containing the human telomeric repeat, has been determined in solution in complex with a fluorinated pentacyclic quino[4,3,2-kl]acridinium cation (RHPS4). RHPS4 has been identified as a potent inhibitor of telomerase at submicromolar levels (IC(50) value of 0.33(+/-0.13)microM), exhibiting a wide differential between telomerase inhibition and acute cellular toxicity. All of the data point to RHPS4 exerting its chemotherapeutic potency through interaction with, and stabilisation of, four-stranded G-quadruplex structures. RHPS4 forms a dynamic interaction with d(TTAGGGT)(4), as evident from 1H and 19F linewidths, with fast exchange between binding sites induced at 318 K. Perturbations to DNA chemical shifts and 24 intermolecular nuclear Overhauser effects (NOEs) identify the 5'-ApG and 5'-GpT steps as the principle intercalation sites; a structural model has been refined using NOE-restrained molecular dynamics. The central G-tetrad core remains intact, with drug molecules stacking at the ends of the G-quadruplex. The partial positive charge on position 13-N of the acridine ring appears to act as a "pseudo" potassium ion and is positioned above the centre of the G-tetrad in the region of high negative charge density. In both ApG and GpT intercalation sites, the drug is seen to converge to the same orientation in which the pi-system of the drug overlaps primarily with two bases of each G-tetrad. The drug is held in place by stacking interactions with the G-tetrads; however, there is some evidence for a more dynamic, weakly stabilised A-tetrad that stacks partially on top of the drug at the 5'-end of the sequence. Together, the interactions of RHPS4 increase the t(m) of the quadruplex by approximately 20 degrees C. There is no evidence for drug intercalation within the G-quadruplex; however, the structural model strongly supports end-stacking interactions with the terminal G-tetrads.
- School of Chemistry, University Park, Nottingham NG7 2RD, UK.
Organizational Affiliation: