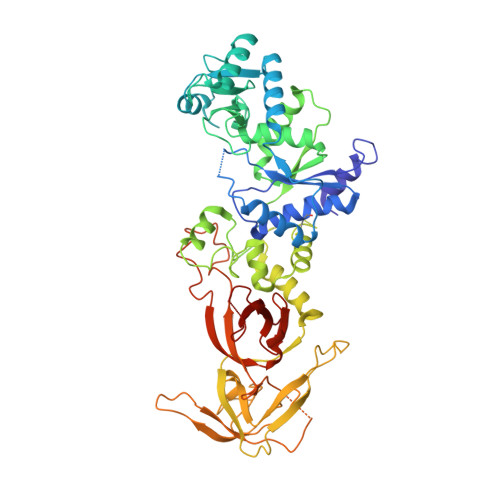
tRNA-Dependent Active Site Assembly in a Class I Aminoacyl-tRNA Synthetase.
Sherlin, L.D., Perona, J.J.(2003) Structure 11: 591-603
- PubMed: 12737824 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00074-1
- Primary Citation Related Structures:
1NYL - PubMed Abstract:
The crystal structure of ligand-free E. coli glutaminyl-tRNA synthetase (GlnRS) at 2.4 A resolution shows that substrate binding is essential to construction of a catalytically proficient active site. tRNA binding generates structural changes throughout the enzyme, repositioning key active site peptides that bind glutamine and ATP. The structure gives insight into longstanding questions regarding the tRNA dependence of glutaminyl adenylate formation, the coupling of amino acid and tRNA selectivities, and the roles of specific pathways for transmission of tRNA binding signals to the active site. Comparative analysis of the unliganded and tRNA-bound structures shows, in detail, how flexibility is built into the enzyme architecture and suggests that the induced-fit transitions are a key underlying determinant of both amino acid and tRNA specificity.
- Department of Chemistry and Biochemistry, University of California, Santa Barbara, Santa Barbara, CA 93106, USA.
Organizational Affiliation: