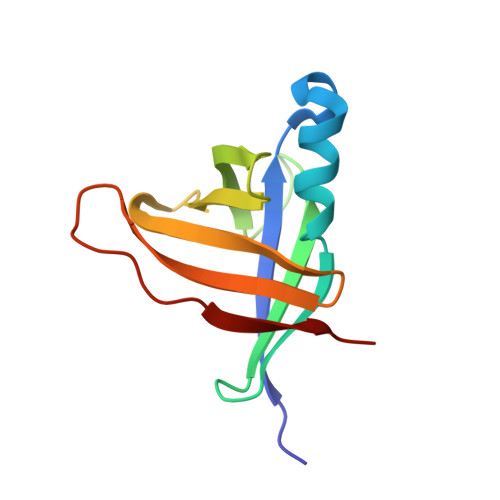
Staphostatins resemble lipocalins, not cystatins in fold.
Rzychon, M., Filipek, R., Sabat, A., Kosowska, K., Dubin, A., Potempa, J., Bochtler, M.(2003) Protein Sci 12: 2252-2256
- PubMed: 14500882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.03247703
- Primary Citation Related Structures:
1NYC - PubMed Abstract:
Staphostatins are the endogenous inhibitors of the major secreted cysteine proteases of Staphylococcus aureus, the staphopains. Here, we present the 1.4 A crystal structure of staphostatin B and show that the fold can be described as a fully closed, highly sheared eight-stranded beta-barrel. Thus, staphostatin B is related to beta-barrel domains that are involved in the inhibition or regulation of proteases of various catalytic types and to the superfamily of lipocalins/cytosolic fatty acid binding proteins. Unexpectedly for a cysteine protease inhibitor, staphostatin B is not significantly similar to cystatins.
- International Institute of Molecular and Cell Biology, 02109 Warsaw, Poland.
Organizational Affiliation: