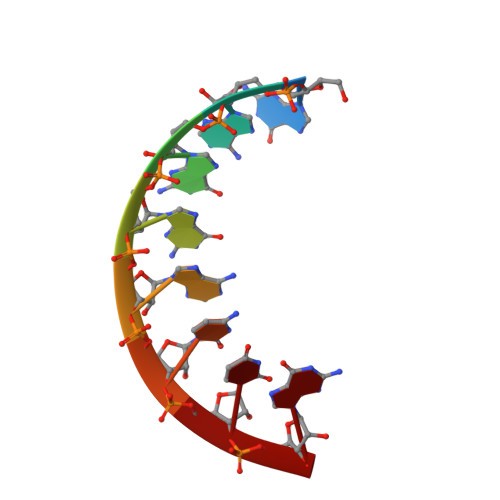
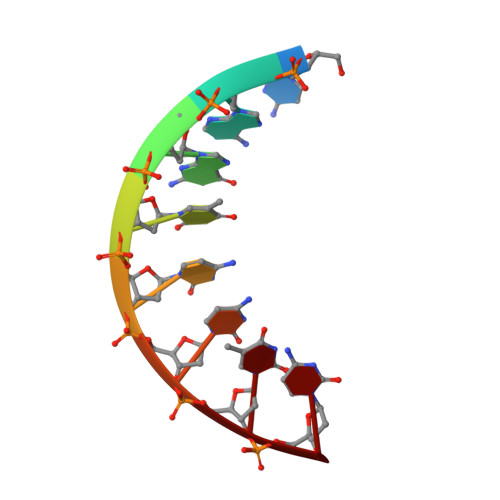
Solution structure of r(gaggacug):d(CAGTCCTC) hybrid: implications for the initiation of HIV-1 (+)-strand synthesis.
Fedoroff, O.Y., Ge, Y., Reid, B.R.(1997) J Mol Biology 269: 225-239
- PubMed: 9191067 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1024
- Primary Citation Related Structures:
1NXR - PubMed Abstract:
The three-dimensional solution structure of the hybrid duplex r(gaggacug):d(CAGTCCTC) has been determined by two-dimensional NMR, distance geometry (DG), restrained molecular dynamics (rMD) and NOE back-calculation methods. This hybrid, consisting of a purine-rich RNA strand and a pyrimidine-rich DNA strand, is related to the polypurine (+)-strand primer formed after (-)-strand DNA synthesis and RNase H degradation of the viral RNA strand and contains the site of a specific cleavage by reverse transcription (RT) RNase H at the end of the HIV-1 polypurine tract. This polypurine primer is an important intermediate in the formation of virally encoded double-stranded DNA prior to HIV-1 retrovirus integration. The correct processing of this primer is vital in the life cycle of the human immunodeficiency virus type (HIV-1) retrovirus. The structure of the r(gaggacug):d(CAGTCCTC) hybrid, as determined in solution by NMR, is intermediate between canonical A-type and B-type double helices, and has mixed structural characteristics. It is quantitatively different from the previously determined solution structures of other RNA-DNA hybrids, particularly in the width and shape of the major groove, which is wider than the major groove of other hybrids and is close to the dimension of the major groove of B-type DNA duplexes. The structure of this hybrid duplex contains a prominent bend in the double helix with a magnitude and direction similar to the bend in Okazaki fragments. The structural features of the present duplex may explain the unique interactions of this sequence with HIV-1 RT during both (-)-strand and (+)-strand DNA synthesis.
- Chemistry Department, University of Washington, Seattle 98195, USA.
Organizational Affiliation: