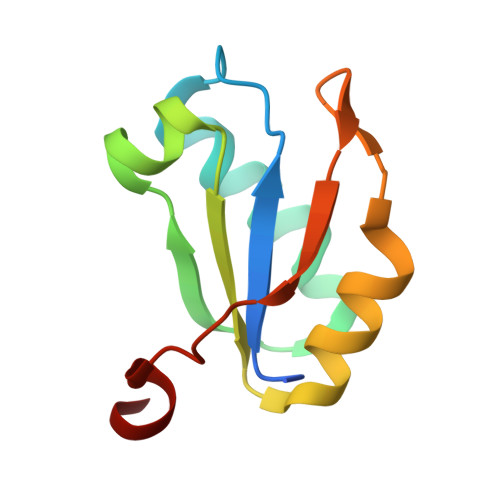
U1A RNA-binding domain at 1.8 A resolution.
Rupert, P.B., Xiao, H., Ferre-D'Amare, A.R.(2003) Acta Crystallogr D Biol Crystallogr 59: 1521-1524
- PubMed: 12876372
- DOI: https://doi.org/10.1107/s0907444903011338
- Primary Citation of Related Structures:
1NU4 - PubMed Abstract:
The human U1A RNA-binding domain (RBD1) adopts one of the most common protein folds, the RNA-recognition motif, and is a paradigm for understanding RNA-protein interactions. A 2.8 A resolution structure of the unbound RBD1 has previously been determined [Nagai et al. (1990). Nature (London), 348, 515-520] and revealed a well defined alpha/beta core with disordered termini. Using a longer construct, a 1.8 A resolution structure of the unbound domain was determined that reveals an ordered C-terminal helix. The presence of this helix is consistent with a solution structure of the free domain [Avis et al. (1996). J. Mol. Biol. 257, 398-411]; however, in the solution structure the helix occludes the RNA-binding surface. In the present structure, the helix occupies a position similar to that seen in a 1.9 A resolution RNA-RBD1 complex structure [Oubridge et al. (1994). Nature (London), 372, 432-438]. The crystals in this study were grown from 2.2 M sodium malonate. It is possible that the high salt concentration helps to orient the C-terminal helix in the RNA-bound conformation by strengthening hydrophobic interactions between the buried face of the helix and the alpha/beta core of the protein. Alternatively, the malonate (several molecules of which are bound in the vicinity of the RNA-binding surface) may mimic RNA.
- Fred Hutchinson Cancer Research Center, USA.
Organizational Affiliation: