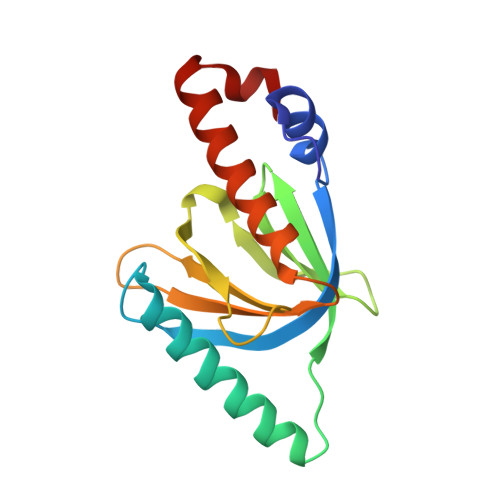
Origins of Peptide Selectivity and Phosphoinositide Binding Revealed by Structures of Disabled-1 PTB Domain Complexes
Stolt, P.C., Jeon, H., Song, H.K., Herz, J., Eck, M.J., Blacklow, S.C.(2003) Structure 11: 569-579
- PubMed: 12737822 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00068-6
- Primary Citation Related Structures:
1NTV, 1NU2 - PubMed Abstract:
Formation of the mammalian six-layered neocortex depends on a signaling pathway that involves Reelin, the very low-density lipoprotein receptor, the apolipoprotein E receptor-2 (ApoER2), and the adaptor protein Disabled-1 (Dab1). The 1.5 A crystal structure of a complex between the Dab1 phosphotyrosine binding (PTB) domain and a 14-residue peptide from the ApoER2 tail explains the unusual preference of Dab1 for unphosphorylated tyrosine within the NPxY motif of the peptide. Crystals of the complex soaked with the phosphoinositide PI-4,5P(2) (PI) show that PI binds to conserved basic residues on the PTB domain opposite the peptide binding groove. This finding explains how the Dab1 PTB domain can simultaneously bind PI and the ApoER2 tail. Recruitment of the Dab1 PTB domain to PI-rich regions of the plasma membrane may facilitate association with the Reelin receptor cytoplasmic tails to transduce a critical positional cue to migrating neurons.
- Department of Pathology, Brigham and Women's Hospital and Harvard Medical School, 75 Francis Street, Boston, MA 02115, USA.
Organizational Affiliation: