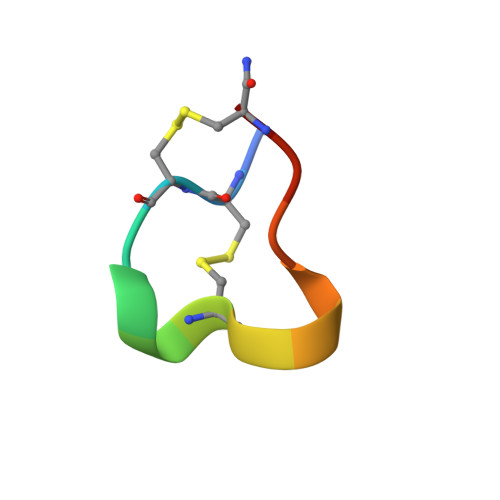
Three-dimensional structure of the alpha-conotoxin GI at 1.2 A resolution
Guddat, L.W., Martin, J.L., Shan, L., Edmundson, A.B., Gray, W.R.(1996) Biochemistry 35: 11329-11335
- PubMed: 8784187 Search on PubMed
- DOI: https://doi.org/10.1021/bi960820h
- Primary Citation Related Structures:
1NOT - PubMed Abstract:
Predatory marine snails of the genus Conus paralyze their fish prey by injecting a potent toxin. The alpha-conotoxin GI is a 13-residue peptide isolated from venom of Conus geographus. It functions by blocking the postsynaptic nicotinic acetylcholine receptor. After crystallization in deionized water, the three-dimensional structure of the GI neurotoxin was determined to 1.2 A resolution by X-ray crystallography. This structure, which can be described as a triangular slab, shows overall similarities to those derived by NMR, CD, and predictive methods. The principal framework of the molecule is provided by two disulfide bonds, one linking Cys 2 and Cys 7 and the other Cys 3 and Cys 13. Opposite ends of the sequence are drawn together even further by hydrogen bonds between Glu 1 and Cys 13 and between Cys 2 and Ser 12. Since the C-terminus is amidated, only one negative charge is present (carboxylate of Glu 1), and this is not implicated in receptor binding. Two positively charged regions (the alpha-amino group of Glu 1 and the guanido group of Arg 9) are situated 15 A apart at the corners of the triangular face of the molecule. phi, psi angles characteristic of a 3(10) helix were observed for residues 5-7. For residues 8-11, these angles were consistent with either a type I beta-turn or a distorted 3(10) helix.
- Centre for Drug Design and Development, University of Queensland, St. Lucia, Australia.
Organizational Affiliation: